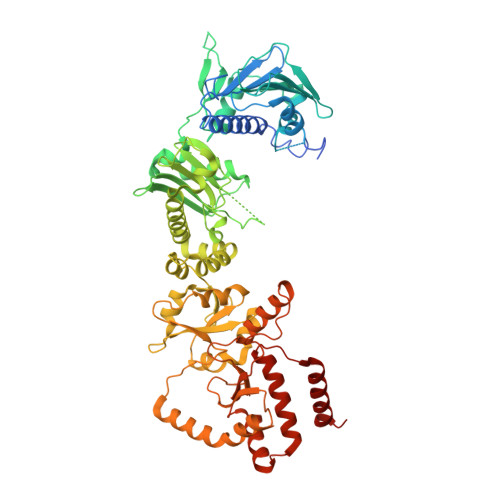
Structures of GRP94-Nucleotide Complexes Reveal Mechanistic Differences between the hsp90 Chaperones.
Dollins, D.E., Warren, J.J., Immormino, R.M., Gewirth, D.T.(2007) Mol Cell 28: 41-56
- PubMed: 17936703
- DOI: https://doi.org/10.1016/j.molcel.2007.08.024
- Primary Citation of Related Structures:
2O1T, 2O1U, 2O1V, 2O1W - PubMed Abstract:
GRP94, an essential endoplasmic reticulum chaperone, is required for the conformational maturation of proteins destined for cell-surface display or export. The extent to which GRP94 and its cytosolic paralog, Hsp90, share a common mechanism remains controversial. GRP94 has not been shown conclusively to hydrolyze ATP or bind cochaperones, and both activities, by contrast, result in conformational changes and N-terminal dimerization in Hsp90 that are critical for its function. Here, we report the 2.4 A crystal structure of mammalian GRP94 in complex with AMPPNP and ADP. The chaperone is conformationally insensitive to the identity of the bound nucleotide, adopting a "twisted V" conformation that precludes N-terminal domain dimerization. We also present conclusive evidence that GRP94 possesses ATPase activity. Our observations provide a structural explanation for GRP94's observed rate of ATP hydrolysis and suggest a model for the role of ATP binding and hydrolysis in the GRP94 chaperone cycle.
- Hauptman-Woodward Medical Research Institute, 700 Ellicott Street, Buffalo, NY 14203, USA.
Organizational Affiliation: