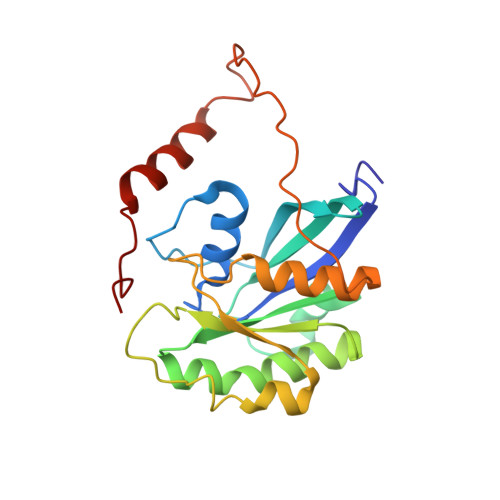
Solution structures of Mengovirus Leader protein, its phosphorylated derivatives, and in complex with nuclear transport regulatory protein, RanGTPase.
Bacot-Davis, V.R., Ciomperlik, J.J., Basta, H.A., Cornilescu, C.C., Palmenberg, A.C.(2014) Proc Natl Acad Sci U S A 111: 15792-15797
- PubMed: 25331866 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1411098111
- Primary Citation Related Structures:
2MMG, 2MMH, 2MMI, 2MMK, 2MML - PubMed Abstract:
Cardiovirus Leader (L) proteins induce potent antihost inhibition of active cellular nucleocytoplasmic trafficking by triggering aberrant hyperphosphorylation of nuclear pore proteins (Nup). To achieve this, L binds protein RanGTPase (Ran), a key trafficking regulator, and diverts it into tertiary or quaternary complexes with required kinases. The activity of L is regulated by two phosphorylation events not required for Ran binding. Matched NMR studies on the unphosphorylated, singly, and doubly phosphorylated variants of Mengovirus L (L(M)) show both modifications act together to partially stabilize a short internal α-helix comprising L(M) residues 43-46. This motif implies that ionic and Van der Waals forces contributed by phosphorylation help organize downstream residues 48-67 into a new interface. The full structure of L(M) as bound to Ran (unlabeled) and Ran (216 aa) as bound by L(M) (unlabeled) places L(M) into the BP1 binding site of Ran, wrapped by the conformational flexible COOH tail. The arrangement explains the tight KD for this complex and places the LM zinc finger and phosphorylation interface as surface exposed and available for subsequent reactions. The core structure of Ran, outside the COOH tail, is not altered by L(M) binding and remains accessible for canonical RanGTP partner interactions. Pull-down assays identify at least one putative Ran:L(M) partner as an exportin, Crm1, or CAS. A model of Ran:L(M):Crm1, based on the new structures suggests LM phosphorylation status may mediate Ran's selection of exportin(s) and cargo(s), perverting these native trafficking elements into the lethal antihost Nup phosphorylation pathways.
- Institute for Molecular Virology and.
Organizational Affiliation: