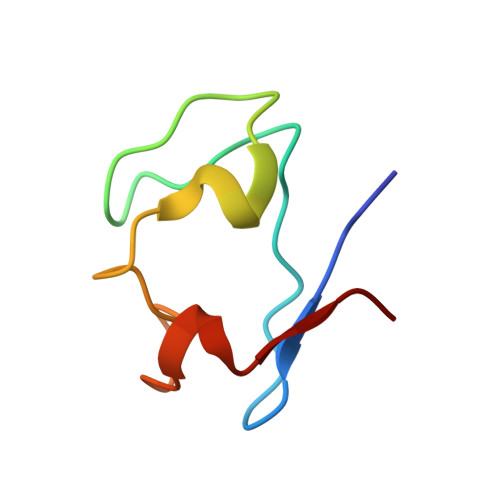
An NMR structural study of nickel-substituted rubredoxin
Goodfellow, B.J., Duarte, I.C., Macedo, A.L., Volkman, B.F., Nunes, S.G., Moura, I., Markley, J.L., Moura, J.J.(2010) J Biol Inorg Chem 15: 409-420
- PubMed: 19997764 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00775-009-0613-6
- Primary Citation Related Structures:
2KKD - PubMed Abstract:
The Ni(II) and Zn(II) derivatives of Desulfovibrio vulgaris rubredoxin (DvRd) have been studied by NMR spectroscopy to probe the structure at the metal centre. The beta CH(2) proton pairs from the cysteines that bind the Ni(II) atom have been identified using 1D nuclear Overhauser enhancement (NOE) difference spectra and sequence specifically assigned via NOE correlations to neighbouring protons and by comparison with the published X-ray crystal structure of a Ni(II) derivative of Clostridium pasteurianum rubredoxin. The solution structures of DvRd(Zn) and DvRd(Ni) have been determined and the paramagnetic form refined using pseudocontact shifts. The determination of the magnetic susceptibility anisotropy tensor allowed the contact and pseudocontact contributions to the observed chemical shifts to be obtained. Analysis of the pseudocontact and contact chemical shifts of the cysteine H beta protons and backbone protons close to the metal centre allowed conclusions to be drawn as to the geometry and hydrogen-bonding pattern at the metal binding site. The importance of NH-S hydrogen bonds at the metal centre for the delocalization of electron spin density is confirmed for rubredoxins and can be extrapolated to metal centres in Cu proteins: amicyanin, plastocyanin, stellacyanin, azurin and pseudoazurin.
- CICECO, Departamento de Química, Universidade Aveiro, 3810-193, Aveiro, Portugal. brian.goodfellow@ua.pt
Organizational Affiliation: