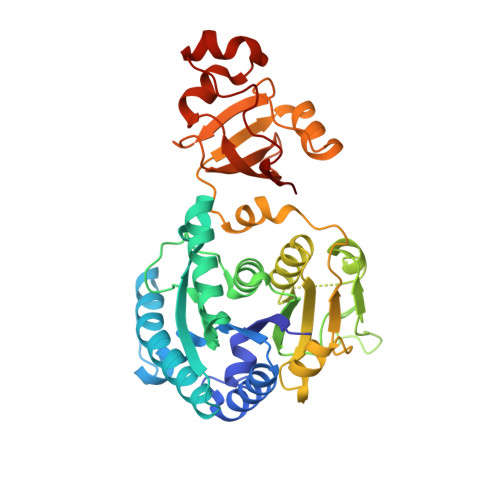
A Novel Two-Domain Architecture within the Amino Acid Kinase Enzyme Family Revealed by the Crystal Structure of Escherichia Coli Glutamate 5-Kinase.
Marco-Marin, C., Gil-Ortiz, F., Perez-Arellano, I., Cervera, J., Fita, I., Rubio, V.(2007) J Mol Biology 367: 1431
- PubMed: 17321544
- DOI: https://doi.org/10.1016/j.jmb.2007.01.073
- Primary Citation Related Structures:
2J5T, 2J5V - PubMed Abstract:
Glutamate 5-kinase (G5K) makes the highly unstable product glutamyl 5-phosphate (G5P) in the initial, controlling step of proline/ornithine synthesis, being feedback-inhibited by proline or ornithine, and causing, when defective, clinical hyperammonaemia. We determined two crystal structures of G5K from Escherichia coli, at 2.9 A and 2.5 A resolution, complexed with glutamate and sulphate, or with G5P, sulphate and the proline analogue 5-oxoproline. E. coli G5K presents a novel tetrameric (dimer of dimers) architecture. Each subunit contains a 257 residue AAK domain, typical of acylphosphate-forming enzymes, with characteristic alpha(3)beta(8)alpha(4) sandwich topology. This domain is responsible for catalysis and proline inhibition, and has a crater on the beta sheet C-edge that hosts the active centre and bound 5-oxoproline. Each subunit contains a 93 residue C-terminal PUA domain, typical of RNA-modifying enzymes, which presents the characteristic beta(5)beta(4) sandwich fold and three alpha helices. The AAK and PUA domains of one subunit associate non-canonically in the dimer with the same domains of the other subunit, leaving a negatively charged hole between them that hosts two Mg ions in one crystal, in line with the G5K requirement for free Mg. The tetramer, formed by two dimers interacting exclusively through their AAK domains, is flat and elongated, and has in each face, pericentrically, two exposed active centres in alternate subunits. This would permit the close apposition of two active centres of bacterial glutamate-5-phosphate reductase (the next enzyme in the proline/ornithine-synthesising route), supporting the postulated channelling of G5P. The structures clarify substrate binding and catalysis, justify the high glutamate specificity, explain the effects of known point mutations, and support the binding of proline near glutamate. Proline binding may trigger the movement of a loop that encircles glutamate, and which participates in a hydrogen bond network connecting active centres, which is possibly involved in the cooperativity for glutamate.
- Instituto de Biomedicina de Valencia (IBV-CSIC) and Center for Biomedical Research on Rare Diseases (CIBERER-ISCIII), Jaume Roig 11, Valencia-46010, Spain.
Organizational Affiliation: