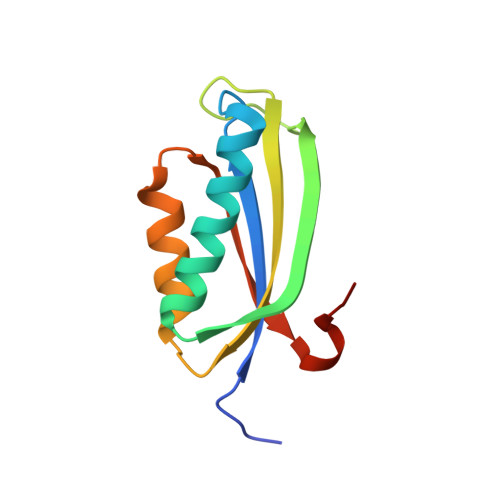
Folding of S6 Structures with Divergent Amino Acid Composition: Pathway Flexibility within Partly Overlapping Foldons.
Olofsson, M., Hansson, S., Hedberg, L., Logan, D.T., Oliveberg, M.(2007) J Mol Biology 365: 237
- PubMed: 17056063 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.09.016
- Primary Citation Related Structures:
2J5A - PubMed Abstract:
Studies of circular permutants have demonstrated that the folding reaction of S6 from Thermus thermophilus (S6(T)) is malleable and responds in an ordered manner to changes of the sequence separation between interacting residues: the S6(T) permutants retain a common nucleation pattern in the form of a two-strand-helix motif that can be recruited from different parts of the structure. To further test the robustness of the two-strand-helix nucleus we have here determined the crystal structure and folding reaction of an evolutionary divergent S6 protein from the hyperthermophilic bacterium Aquifex aeolicus (S6(A)). Although the overall topology of S6(A) is very similar to that of S6(T) the architecture of the hydrophobic core is radically different by containing a large proportion of stacked Phe side-chains. Despite this disparate core composition, the folding rate constant and the kinetic m values of S6(A) are identical to those of S6(T). The folding nucleus of S6(A) is also found to retain the characteristic two-strand-helix motif of the S6(T) permutants, but with a new structural emphasis. The results suggest that the protein folding reaction is linked to topology only in the sense that the native-state topology determines the repertoire of accessible nucleation motifs. If the native structure allows several equivalent ways of recruiting a productive nucleus the folding reaction is free to redistribute within these topological constraints.
- Department of Biochemistry, Umeå University, SE-901 87 Umeå, Sweden.
Organizational Affiliation: