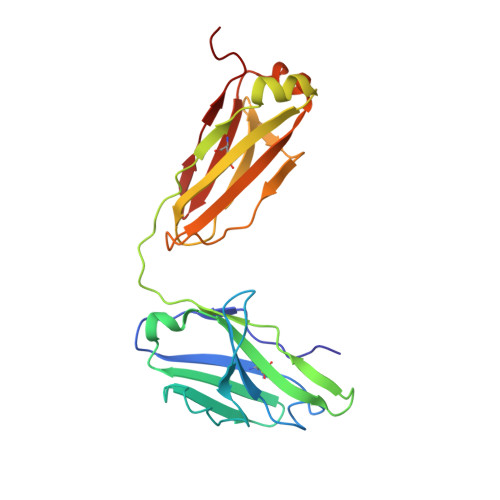
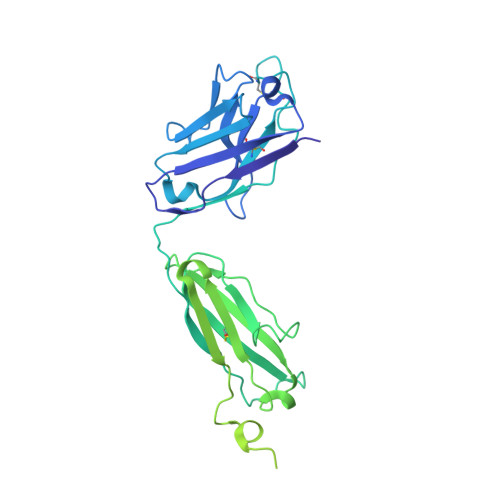
The primary structure of crystallizable monoclonal immunoglobulin IgG1 Kol. II. Amino acid sequence of the L-chain, gamma-type, subgroup I
Kratzin, H.D., Palm, W., Stangel, M., Schmidt, W.E., Friedrich, J., Hilschmann, N.(1989) Biol Chem Hoppe Seyler 370: 263-272
- PubMed: 2713105 Search on PubMed
- Primary Citation Related Structures:
2FB4, 2IG2 - PubMed Abstract:
The immunoglobulin Kol was the first intact antibody molecule which was characterized by high-resolution X-ray crystallography. Furthermore the complete amino-acid sequence of the heavy (H)-chain is known. Here we report the complete amino-acid sequence of the light (L)-chain of the monoclonal immunoglobulin Kol (IgG1). The polypeptide has an Mr of 22,781, consists of 216 amino acids and due to its structure is of the lambda-type. With the characteristic amino acids threonine, asparagine, threonine, glycine and lysine in positions 101, 114, 116, 154, and 165, respectively the Kol L-chain is of the Mcg isotype. With the proteins Mcg, Mot, Bur, Loc and Mem six myeloma-derived amino-acid sequences of the same isotype are known. The amino-acid sequence of the N-terminal variable part is characteristic of subgroup 1. This contribution completes the primary structure of IgG1 Kol.
- max-Planck-Institut für experimentelle Medizin, Abteilung Immunchemie, Göttingen.
Organizational Affiliation: