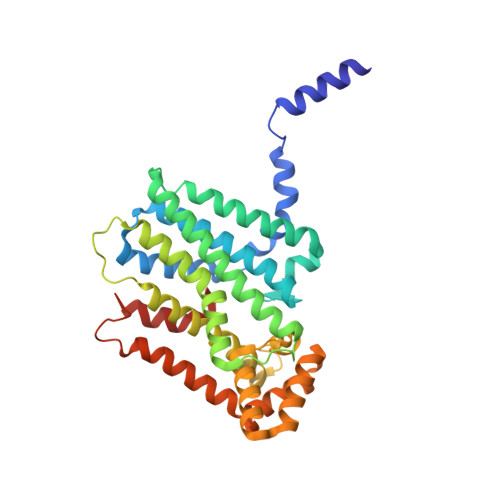
Crystal Structure of Type-III Geranylgeranyl Pyrophosphate Synthase from Saccharomyces cerevisiae and the Mechanism of Product Chain Length Determination.
Chang, T.-H., Guo, R.-T., Ko, T.-P., Wang, A.H., Liang, P.-H.(2006) J Biological Chem 281: 14991-15000
- PubMed: 16554305 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M512886200
- Primary Citation Related Structures:
2DH4 - PubMed Abstract:
Geranylgeranyl pyrophosphate synthase (GGPPs) catalyzes a condensation reaction of farnesyl pyrophosphate with isopentenyl pyrophosphate to generate C(20) geranylgeranyl pyrophosphate, which is a precursor for carotenoids, chlorophylls, geranylgeranylated proteins, and archaeal ether-linked lipid. For short-chain trans-prenyltransferases that synthesize C(10)-C(25) products, bulky amino acid residues generally occupy the fourth or fifth position upstream from the first DDXXD motif to block further elongation of the final products. However, the short-chain type-III GGPPs in eukaryotes lack any large amino acid at these positions. In this study, the first structure of type-III GGPPs from Saccharomyces cerevisiae has been determined to 1.98 A resolution. The structure is composed entirely of 15 alpha-helices joined by connecting loops and is arranged with alpha-helices around a large central cavity. Distinct from other known structures of trans-prenyltransferases, the N-terminal 17 amino acids (9-amino acid helix A and the following loop) of this GGPPs protrude from the helix core into the other subunit and contribute to the tight dimer formation. Deletion of the first 9 or 17 amino acids caused the dissociation of dimer into monomer, and the Delta(1-17) mutant showed abolished enzyme activity. In each subunit, an elongated hydrophobic crevice surrounded by D, F, G, H, and I alpha-helices contains two DDXXD motifs at the top for substrate binding with one Mg(2+) coordinated by Asp(75), Asp(79), and four water molecules. It is sealed at the bottom with three large residues of Tyr(107), Phe(108), and His(139). Compared with the major product C(30) synthesized by mutant H139A, the products generated by mutant Y107A and F108A are predominantly C(40) and C(30), respectively, suggesting the most important role of Tyr(107) in determining the product chain length.
- Institute of Biochemical Sciences, National Taiwan University, Taipei 106, Taiwan.
Organizational Affiliation: