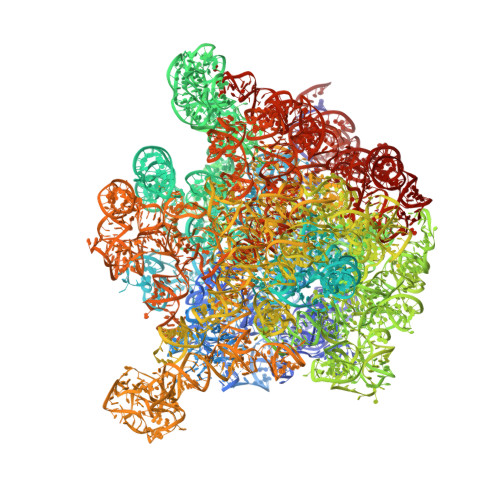
The Binding Mode of the Trigger Factor on the Ribosome: Implications for Protein Folding and SRP Interaction
Schlunzen, F., Wilson, D.N., Tian, P., Harms, J.M., McInnes, S.J., Hansen, H.A., Albrecht, R., Buerger, J., Wilbanks, S.M., Fucini, P.(2005) Structure 13: 1685-1694
- PubMed: 16271892 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.08.007
- Primary Citation Related Structures:
2D3O - PubMed Abstract:
This study presents the X-ray structure of the N-terminal binding domain of the D. radiodurans trigger factor (TF) in complex with the D. radiodurans large ribosomal subunit. At 3.35 A, a complete description of the interactions with ribosomal proteins L23, L29, and 23S rRNA are disclosed, many of which differ from those found previously for a heterologous bacterial-archaeal TF-ribosome complex. The beta hairpin loop of eubacterial L24, which is shorter in archaeal ribosomes, contacts the TF and severely diminishes the molecular cradle proposed to exist between the TF and ribosome. Bound to the ribosome, TF exposes a hydrophobic crevice large enough to accommodate the nascent polypeptide chain. Superimposition of the full-length TF and the signal-recognition particle (SRP) onto the complex shows that simultaneous cohabitation is possible, in agreement with biochemical data, and suggests a model for the interplay of TF, SRP, and the nascent chain during translation.
- Max-Planck Institute for Molecular Genetics, Ihnestrasse 73, D-14195 Berlin, Germany. Schluenz@molgen.mpg.de
Organizational Affiliation: