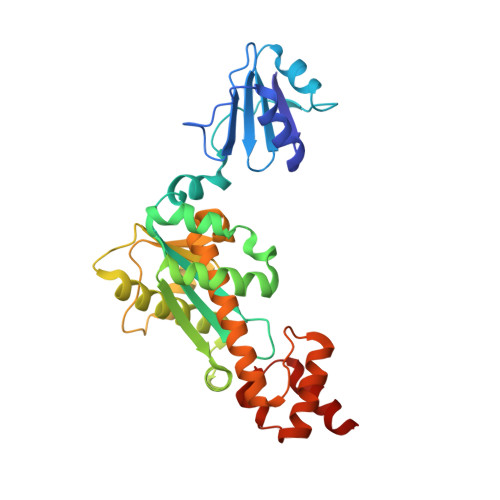
Crystal structure and structure-based mutational analyses of RNase HIII from Bacillus stearothermophilus: a new type 2 RNase H with TBP-like substrate-binding domain at the N terminus
Chon, H., Matsumura, H., Koga, Y., Takano, K., Kanaya, S.(2006) J Mol Biology 356: 165-178
- PubMed: 16343535
- DOI: https://doi.org/10.1016/j.jmb.2005.11.017
- Primary Citation Related Structures:
2D0A, 2D0B, 2D0C - PubMed Abstract:
Ribonuclease HIII (Bst-RNase HIII) from the moderate thermophile Bacillus stearothermophilus is a type 2 RNase H but shows poor amino acid sequence identity with another type 2 RNase H, RNase HII. It is composed of 310 amino acid residues and acts as a monomer. Bst-RNase HIII has a large N-terminal extension with unknown function and a unique active-site motif (DEDE), both of which are characteristics common to RNases HIII. To understand the role of these N-terminal extension and active-site residues, the crystal structure of Bst-RNase HIII was determined in both metal-free and metal-bound forms at 2.1-2.6 angstroms resolutions. According to these structures, Bst-RNase HIII consists of the N-terminal domain and C-terminal RNase H domain. The structures of the N and C-terminal domains were similar to those of TATA-box binding proteins and archaeal RNases HII, respectively. The steric configurations of the four conserved active-site residues were very similar to those of other type 1 and type 2 RNases H. Single Mn and Mg ions were coordinated with Asp97, Glu98, and Asp202, which correspond to Asp10, Glu48, and Asp70 of Escherichia coli RNase HI, respectively. The mutational studies indicated that the replacement of either one of these residues with Ala resulted in a great reduction of the enzymatic activity. Overproduction, purification, and characterization of the Bst-RNase HIII derivatives with N and/or C-terminal truncations indicated that the N-terminal domain and C-terminal helix are involved in substrate binding, but the former contributes to substrate binding more greatly than the latter.
- Department of Material and Life Science, Graduate School of Engineering, Osaka University, 2-1 Yamadaoka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: