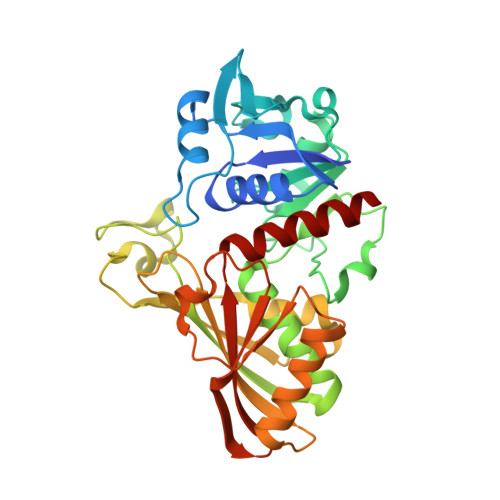
Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 A resolution reveals intriguing extra electron density in the active site
Robien, M.A., Bosch, J., Buckner, F.S., Van Voorhis, W.C., Worthey, E.A., Myler, P., Mehlin, C., Boni, E.E., Kalyuzhniy, O., Anderson, L., Lauricella, A., Gulde, S., Luft, J.R., Detitta, G., Caruthers, J.M., Hodgson, K.O., Soltis, M., Zucker, F., Verlinde, C.L., Merritt, E.A., Schoenfeld, L.W., Hol, W.G.(2006) Proteins 62: 570-577
- PubMed: 16345073 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20801
- Primary Citation Related Structures:
2B4R, 2B4T - PubMed Abstract:
The crystal structure of D-glyceraldehyde-3-phosphate dehydrogenase (PfGAPDH) from the major malaria parasite Plasmodium falciparum is solved at 2.25 A resolution. The structure of PfGAPDH is of interest due to the dependence of the malaria parasite in infected human erythrocytes on the glycolytic pathway for its energy generation. Recent evidence suggests that PfGAPDH may also be required for other critical activities such as apical complex formation. The cofactor NAD(+) is bound to all four subunits of the tetrameric enzyme displaying excellent electron densities. In addition, in all four subunits a completely unexpected large island of extra electron density in the active site is observed, approaching closely the nicotinamide ribose of the NAD(+). This density is most likely the protease inhibitor AEBSF, found in maps from two different crystals. This putative AEBSF molecule is positioned in a crucial location and hence our structure, with expected and unexpected ligands bound, can be of assistance in lead development and design of novel antimalarials.
- Structural Genomics of Pathogenic Protozoa (SGPP), Department of Biochemistry, University of Washington, Seattle 98195, USA.
Organizational Affiliation: