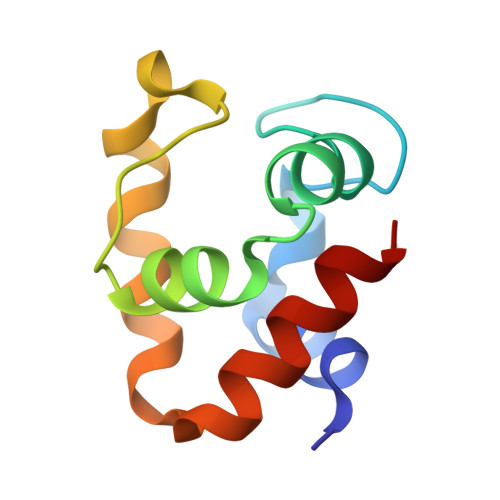
Hyperstability and crystal structure of cytochrome c(555) from hyperthermophilic Aquifex aeolicus
Obuchi, M., Kawahara, K., Motooka, D., Nakamura, S., Yamanaka, M., Takeda, T., Uchiyama, S., Kobayashi, Y., Ohkubo, T., Sambongi, Y.(2009) Acta Crystallogr D Biol Crystallogr 65: 804-813
- PubMed: 19622864 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444909017314
- Primary Citation Related Structures:
2ZXY - PubMed Abstract:
In order to elucidate the relationship between the stability and the structure of the monohaem cytochrome c(555) (AA c(555)) from the hyperthermophilic bacterium Aquifex aeolicus, chemical denaturation and crystal structure determination were carried out. AA c(555) exhibited higher stability than the thermophilic Hydrogenobacter thermophilus cytochrome c(552) (HT c(552)), which is one of the most stable cytochromes c. The three-dimensional crystal structure of AA c(555), which was determined using the multiple anomalous dispersion technique at 1.15 A resolution, included a unique 14-residue extra helix, while the side-chain interactions of several amino-acid residues responsible for the stability of HT c(552) were conserved in AA c(555). The side chain of the Met61 residue in the extra helix was aligned towards the haem, forming a coordination bond between the Met S and haem Fe atoms. In other cytochromes c the corresponding regions always form Omega loops which also include the haem-liganding Met residue and are known to be involved in the initial step in cytochrome c denaturation. The formation of the extra helix in AA c(555) results in the highest helix content, 59.8%, among the monohaem cytochromes c. The extra helix should mainly contribute to the hyperstability of AA c(555) and is presumed to be a novel strategy of cytochromes c for adaptation to a hyperthermophilic environment.
- Hiroshima University, Japan.
Organizational Affiliation: