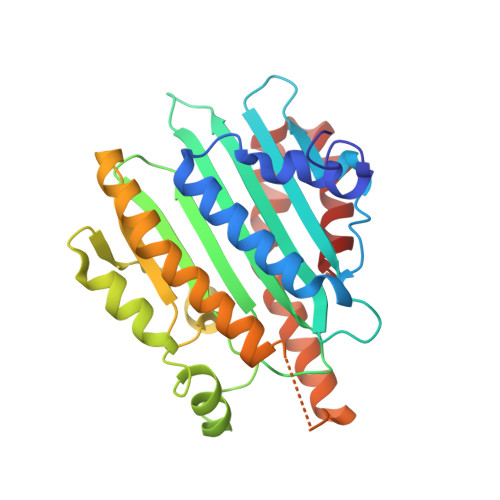
Crystal structure of red chlorophyll catabolite reductase: enlargement of the ferredoxin-dependent bilin reductase family
Sugishima, M., Kitamori, Y., Noguchi, M., Kohchi, T., Fukuyama, K.(2009) J Mol Biology 389: 376-387
- PubMed: 19374909 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.04.017
- Primary Citation Related Structures:
2ZXK, 2ZXL - PubMed Abstract:
The key steps in the degradation pathway of chlorophylls are the ring-opening reaction catalyzed by pheophorbide a oxygenase and sequential reduction by red chlorophyll catabolite reductase (RCCR). During these steps, chlorophyll catabolites lose their color and phototoxicity. RCCR catalyzes the ferredoxin-dependent reduction of the C20/C1 double bond of red chlorophyll catabolite. RCCR appears to be evolutionarily related to the ferredoxin-dependent bilin reductase (FDBR) family, which synthesizes a variety of phytobilin pigments, on the basis of sequence similarity, ferredoxin dependency, and the common tetrapyrrole skeleton of their substrates. The evidence, however, is not robust; the identity between RCCR and FDBR HY2 from Arabidopsis thaliana is only 15%, and the oligomeric states of these enzymes are different. Here, we report the crystal structure of A. thaliana RCCR at 2.4 A resolution. RCCR forms a homodimer, in which each subunit folds in an alpha/beta/alpha sandwich. The tertiary structure of RCCR is similar to those of FDBRs, strongly supporting that these enzymes evolved from a common ancestor. The two subunits are related by noncrystallographic 2-fold symmetry in which the alpha-helices near the edge of the beta-sheet unique in RCCR participate in intersubunit interaction. The putative RCC-binding site, which was derived by superimposing RCCR onto biliverdin-bound forms of FDBRs, forms an open pocket surrounded by conserved residues among RCCRs. Glu154 and Asp291 of A. thaliana RCCR, which stand opposite each other in the pocket, likely are involved in substrate binding and/or catalysis.
- Department of Medical Biochemistry, Kurume University School of Medicine, Fukuoka 830-0011, Japan.
Organizational Affiliation: