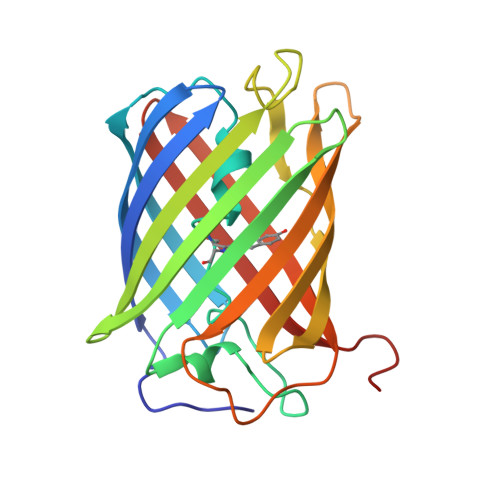
Crystal structure of a new cyan fluorescent protein and its hue-shifted variants
Kikuchi, A., Fukumura, E., Karasawa, S., Shiro, Y., Miyawaki, A.(2009) Biochemistry 48: 5276-5283
- PubMed: 19402703 Search on PubMed
- DOI: https://doi.org/10.1021/bi801658p
- Primary Citation Related Structures:
2ZO6, 2ZO7 - PubMed Abstract:
Green fluorescent protein (GFP) based techniques are well established in molecular biology; however, the detailed mechanism for the fine-tuning of fluorescent colors remains unclear. Here, we report the cloning and crystal structure of a new cyan-emitting GFP-like protein, KCy. We also developed a mutant protein with a high folding efficiency (KCy-G4219: lambda(abs) = 453 nm; lambda(em) = 486 nm). X-ray diffraction analysis revealed that the KCy chromophore is formed from an internal Ser62-Tyr63-Gly64 tripeptide. The serine residue at the first position of the chromophore-forming tripeptide has a short polar chain (-OH) that forms a noncovalent interaction with the His38 imidazole at a distance of 2.96 A. Substitution of His38 in KCy-G4219 with Gln (KCy-R1) or Leu residues resulted in a slight but significant red shift of the emission peak maximum from 486 to 492 or 496 nm, respectively. The crystal structure of KCy-R1 determined at a resolution of 1.58 A showed that the noncovalent interaction between Ser62-OH and the substituted Gln38 occurred over a longer distance (3.07 A) than that observed in the wild-type KCy. Such an interaction is absent in the Leu mutant, suggesting that this interaction is one of the key factors responsible for fine-tuning the emission peak maxima, which are affected by chromophore polarization. Moreover, the structural comparison suggests that an additional water molecule buried in the space between the Ala158 residue and the chromophore phenolate is also responsible for the chromophore polarization.
- Biometal Science Laboratory, RIKEN SPring-8 Center, 1-1-1, Kouto, Sayo, Hyogo 679-5148, Japan. kikuchi@spring8.or.jp
Organizational Affiliation: