Identification of a key element for hydrogen-bonding patterns between protein kinases and their inhibitors.
Katayama, N., Orita, M., Yamaguchi, T., Hisamichi, H., Kuromitsu, S., Kurihara, H., Sakashita, H., Matsumoto, Y., Fujita, S., Niimi, T.(2008) Proteins 73: 795-801
- PubMed: 18767165 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22207
- Primary Citation Related Structures:
2Z7L, 2Z8C - PubMed Abstract:
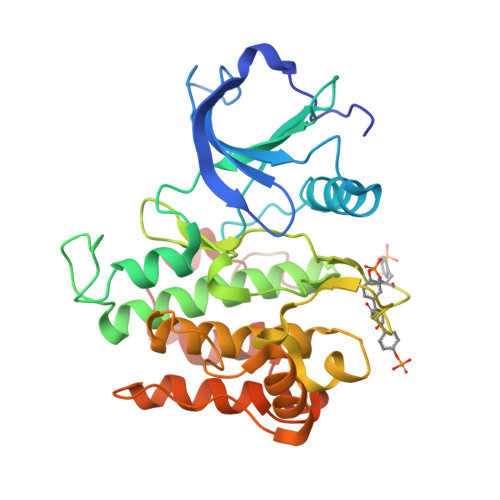
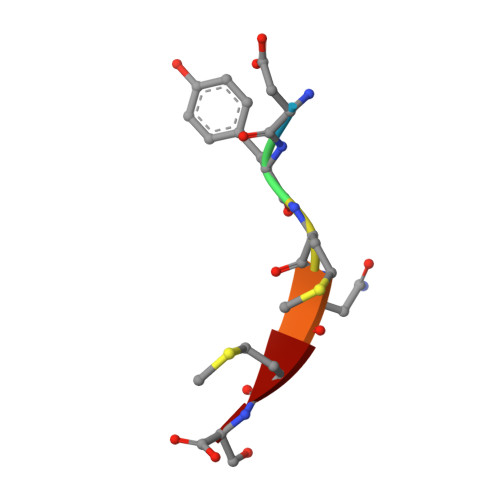
In this article, we report crystal structures for inhibitor-kinase complexes in which the inhibitor has different binding orientations and hydrogen-bonding patterns with extracellular-signal regulated kinase 2 and insulin receptor tyrosine kinase. Our crystallographic studies, and sequence and structural analyses of 532 coordinates of kinases held in the Protein Data Bank, suggest that the length of the "specificity linker" described here is a key structural element of the hydrogen-bonding patterns between protein kinases and their inhibitors.
- Drug Discovery Research, Astellas Pharma Inc., 21 Miyukigaoka, Tsukuba, Ibaraki 305-8585, Japan.
Organizational Affiliation: