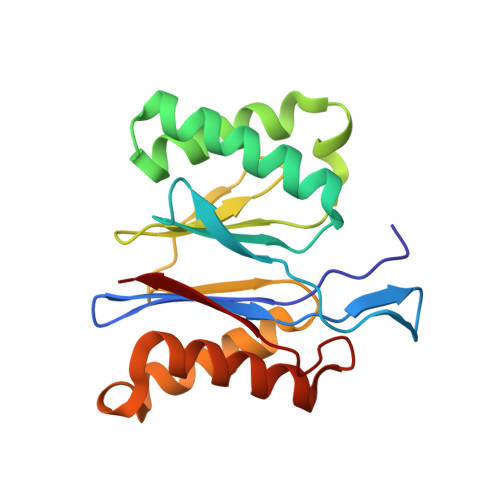
Crystal structure of Bacillus subtilis CodW, a noncanonical HslV-like peptidase with an impaired catalytic apparatus
Rho, S.H., Park, H.H., Kang, G.B., Im, Y.J., Kang, M.S., Lim, B.K., Seong, I.S., Seol, J., Chung, C.H., Wang, J., Eom, S.H.(2007) Proteins 71: 1020-1026
- PubMed: 17979190 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21758
- Primary Citation Related Structures:
2Z3A, 2Z3B - Department of Life Science, Cell Dynamics Research Center, Gwangju Institute of Science and Technology, Gwangju 500-712, Korea.
Organizational Affiliation: