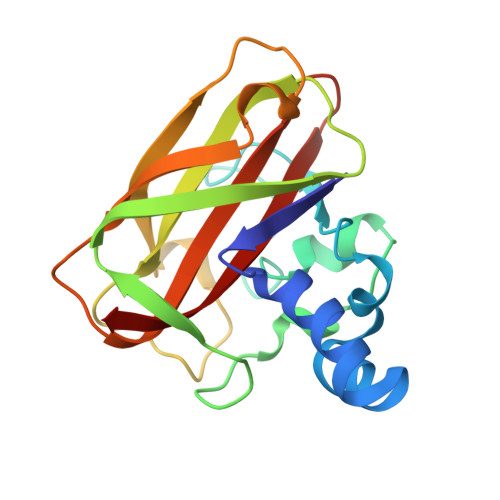
The Copper Active Site of Cbm33 Polysaccharide Oxygenases.
Hemsworth, G.R., Taylor, E.J., Kim, R.Q., Gregory, R.C., Lewis, S.J., Turkenburg, J.P., Parkin, A., Davies, G.J., Walton, P.H.(2013) J Am Chem Soc 135: 6069
- PubMed: 23540833 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja402106e
- Primary Citation Related Structures:
2YOW, 2YOX, 2YOY - PubMed Abstract:
The capacity of metal-dependent fungal and bacterial polysaccharide oxygenases, termed GH61 and CBM33, respectively, to potentiate the enzymatic degradation of cellulose opens new possibilities for the conversion of recalcitrant biomass to biofuels. GH61s have already been shown to be unique metalloenzymes containing an active site with a mononuclear copper ion coordinated by two histidines, one of which is an unusual τ-N-methylated N-terminal histidine. We now report the structural and spectroscopic characterization of the corresponding copper CBM33 enzymes. CBM33 binds copper with high affinity at a mononuclear site, significantly stabilizing the enzyme. X-band EPR spectroscopy of Cu(II)-CBM33 shows a mononuclear type 2 copper site with the copper ion in a distorted axial coordination sphere, into which azide will coordinate as evidenced by the concomitant formation of a new absorption band in the UV/vis spectrum at 390 nm. The enzyme's three-dimensional structure contains copper, which has been photoreduced to Cu(I) by the incident X-rays, confirmed by X-ray absorption/fluorescence studies of both aqueous solution and intact crystals of Cu-CBM33. The single copper(I) ion is ligated in a T-shaped configuration by three nitrogen atoms from two histidine side chains and the amino terminus, similar to the endogenous copper coordination geometry found in fungal GH61.
- Department of Chemistry, University of York, Heslington, York YO10 5DD, United Kingdom.
Organizational Affiliation: