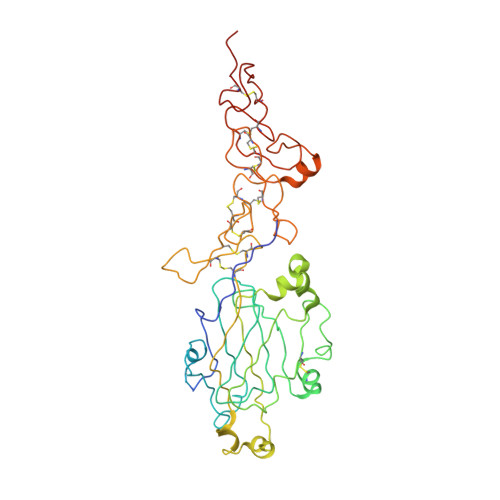
Determinants of Laminin Polymerisation Revealed by the Crystal Structure of the Alpha5 Chain Amino-Terminal Region
Hussain, S.A., Carafoli, F., Hohenester, E.(2011) EMBO Rep 12: 276
- PubMed: 21311558 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2011.3
- Primary Citation Related Structures:
2Y38 - PubMed Abstract:
The polymerization of laminin into a cell-associated network--a key step in basement membrane assembly--is mediated by the laminin amino-terminal (LN) domains at the tips of the three short arms of the laminin αβγ-heterotrimer. The crystal structure of a laminin α5LN-LE1-2 fragment shows that the LN domain is a β-jelly roll with several elaborate insertions that is attached like a flower head to the stalk-like laminin-type epidermal growth factor-like tandem. A surface loop that is strictly conserved in the LN domains of all α-short arms is required for stable ternary association with the β- and γ-short arms in the laminin network.
- Department of Life Sciences, Biophysics Section, Blackett Laboratory, Prince Consort Road, Imperial College London, London SW7 2AZ, UK.
Organizational Affiliation: