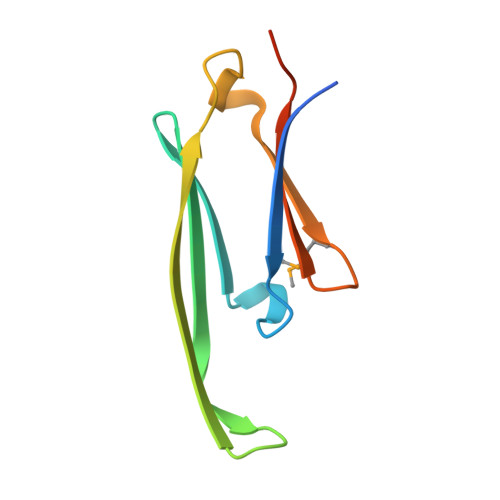
Crystal Structure of R120G Disease Mutant of Human Alphab-Crystallin Domain Dimer Shows Closure of a Groove
Clark, A.R., Naylor, C.E., Bagneris, C., Keep, N.H., Slingsby, C.(2011) J Mol Biology 408: 118
- PubMed: 21329698 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.02.020
- Primary Citation Related Structures:
2Y1Y, 2Y1Z, 2Y22 - PubMed Abstract:
Small heat shock proteins form large cytosolic assemblies from an "α-crystallin domain" (ACD) flanked by sequence extensions. Mutation of a conserved arginine in the ACD of several human small heat shock protein family members causes many common inherited diseases of the lens and neuromuscular system. The mutation R120G in αB-crystallin causes myopathy, cardiomyopathy and cataract. We have solved the X-ray structure of the excised ACD dimer of human αB R120G close to physiological pH and compared it with several recently determined wild-type vertebrate ACD dimer structures. Wild-type excised ACD dimers have a deep groove at the interface floored by a flat extended "bottom sheet." Solid-state NMR studies of large assemblies of full-length αB-crystallin have shown that the groove is blocked in the ACD dimer by curvature of the bottom sheet. The crystal structure of R120G ACD dimer also reveals a closed groove, but here the bottom sheet is flat. Loss of Arg120 results in rearrangement of an extensive array of charged interactions across this interface. His83 and Asp80 on movable arches on either side of the interface close the groove by forming two new salt bridges. The residues involved in this extended set of ionic interactions are conserved in Hsp27, Hsp20, αA- and αB-crystallin sequences. They are not conserved in Hsp22, where mutation of the equivalent of Arg120 causes neuropathy. We speculate that the αB R120G mutation disturbs oligomer dynamics, causing the growth of large soluble oligomers that are toxic to cells by blocking essential processes.
- Department of Biological Sciences, Crystallography, Institute of Structural and Molecular Biology, Birkbeck College, Malet Street, London WC1E 7HX, UK.
Organizational Affiliation: