Structure and Properties of a Complex of Alpha-Synuclein and a Single-Domain Camelid Antibody.
De Genst, E.J., Guilliams, T., Wellens, J., O'Day, E.M., Waudby, C.A., Meehan, S., Dumoulin, M., Hsu, S.-T.D., Cremades, N., Verschueren, K.H.G., Pardon, E., Wyns, L., Steyaert, J., Christodoulou, J., Dobson, C.M.(2010) J Mol Biology 402: 326
- PubMed: 20620148 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.07.001
- Primary Citation Related Structures:
2X6M - PubMed Abstract:
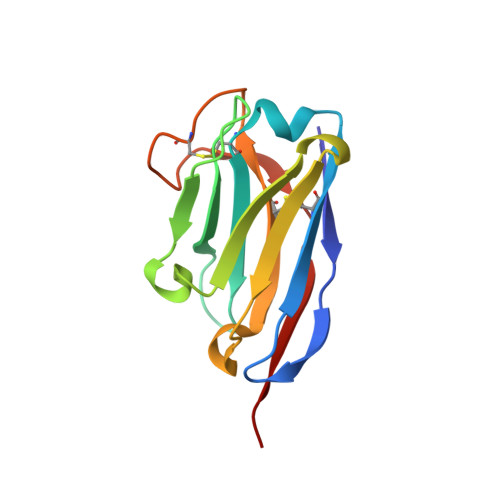
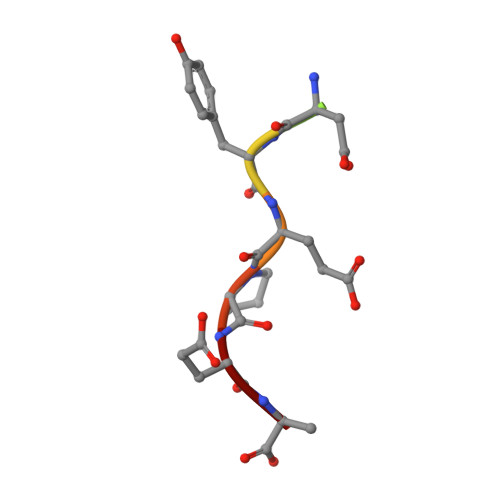
The aggregation of the intrinsically disordered protein α-synuclein to form fibrillar amyloid structures is intimately associated with a variety of neurological disorders, most notably Parkinson's disease. The molecular mechanism of α-synuclein aggregation and toxicity is not yet understood in any detail, not least because of the paucity of structural probes through which to study the behavior of such a disordered system. Here, we describe an investigation involving a single-domain camelid antibody, NbSyn2, selected by phage display techniques to bind to α-synuclein, including the exploration of its effects on the in vitro aggregation of the protein under a variety of conditions. We show using isothermal calorimetric methods that NbSyn2 binds specifically to monomeric α-synuclein with nanomolar affinity and by means of NMR spectroscopy that it interacts with the four C-terminal residues of the protein. This latter finding is confirmed by the determination of a crystal structure of NbSyn2 bound to a peptide encompassing the nine C-terminal residues of α-synuclein. The NbSyn2:α-synuclein interaction is mediated mainly by side-chain interactions while water molecules cross-link the main-chain atoms of α-synuclein to atoms of NbSyn2, a feature we believe could be important in intrinsically disordered protein interactions more generally. The aggregation behavior of α-synuclein at physiological pH, including the morphology of the resulting fibrillar structures, is remarkably unaffected by the presence of NbSyn2 and indeed we show that NbSyn2 binds strongly to the aggregated as well as to the soluble forms of α-synuclein. These results give strong support to the conjecture that the C-terminal region of the protein is not directly involved in the mechanism of aggregation and suggest that binding of NbSyn2 could be a useful probe for the identification of α-synuclein aggregation in vitro and possibly in vivo.
- Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB2 1EW, UK.
Organizational Affiliation: