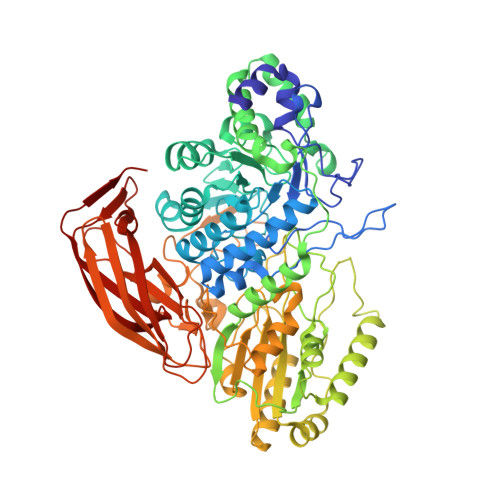
Structural and Functional Analysis of Beta-Glucosidase 3B from Thermotoga Neapolitana: A Thermostable 3-Domain Representative of Glycoside Hydrolase Family 3
Pozzo, T., Linares Pasten, J., Karlsson, E.N., Logan, D.T.(2010) J Mol Biology 397: 724
- PubMed: 20138890 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.01.072
- Primary Citation Related Structures:
2X40, 2X41, 2X42 - PubMed Abstract:
Based on sequence and phylogenetic analyses, glycoside hydrolase (GH) family 3 can be divided into several clusters that differ in the length of their primary sequences. However, structural data on representatives of GH3 are still scarce, since only three of their structures are known and only one of them has been thoroughly characterized-that of an exohydrolase from barley. To allow a deeper structural understanding of the GH3 family, we have determined the crystal structure of the thermostable beta-glucosidase from Thermotoga neapolitana, which has potentially important applications in environmentally friendly industrial biosynthesis at a resolution of 2.05 A. Selected active-site mutants have been characterized kinetically, and the structure of the mutant D242A is presented at 2.1 A resolution. Bgl3B from Th. neapolitana is the first example of a GH3 glucosidase with a three-domain structure. It is composed of an (alpha/beta)(8) domain similar to a triose phosphate isomerase barrel, a five-stranded alpha/beta sandwich domain (both of which are important for active-site organization), and a C-terminal fibronectin type III domain of unknown function. Remarkably, the direction of the second beta-strand of the triose phosphate isomerase barrel domain is reversed, which has implications for the active-site shape. The active site, at the interface of domains 1 and 2, is much more open to solvent than the corresponding site in the structurally homologous enzyme from barley, and only the -1 site is well defined. The structures, in combination with kinetic studies of active-site variants, allow the identification of essential catalytic residues (the nucleophile D242 and the acid/base E458), as well as other residues at the -1 subsite, including D58 and W243, which, by mutagenesis, are shown to be important for substrate accommodation/interaction. The position of the fibronectin type III domain excludes a direct participation of this domain in the recognition of small substrates, although it may be involved in the anchoring of the enzyme on large polymeric substrates and in thermostability.
- Department of Biotechnology, Center for Chemistry and Chemical Engineering, Lund University, Lund, Sweden.
Organizational Affiliation: