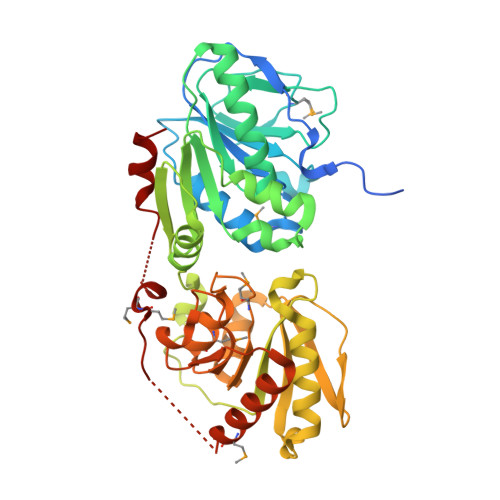
Structural Basis of Substrate Binding in Wsaf, a Rhamnosyltransferase from Geobacillus Stearothermophilus.
Steiner, K., Hagelueken, G., Messner, P., Schaeffer, C., Naismith, J.H.(2010) J Mol Biology 397: 436
- PubMed: 20097205 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.01.035
- Primary Citation Related Structures:
2X0D, 2X0E, 2X0F - PubMed Abstract:
Carbohydrate polymers are medically and industrially important. The S-layer of many Gram-positive organisms comprises protein and carbohydrate polymers and forms an almost paracrystalline array on the cell surface. Not only is this array important for the bacteria but it has potential application in the manufacture of commercially important polysaccharides and glycoconjugates as well. The S-layer glycoprotein glycan from Geobacillus stearothermophilus NRS 2004/3a is mainly composed of repeating units of three rhamnose sugars linked by alpha-1,3-, alpha-1,2-, and beta-1,2-linkages. The formation of the beta-1,2-linkage is catalysed by the enzyme WsaF. The rational use of this system is hampered by the fact that WsaF and other enzymes in the pathway share very little homology to other enzymes. We report the structural and biochemical characterisation of WsaF, the first such rhamnosyltransferase to be characterised. Structural work was aided by the surface entropy reduction method. The enzyme has two domains, the N-terminal domain, which binds the acceptor (the growing rhamnan chain), and the C-terminal domain, which binds the substrate (dTDP-beta-l-rhamnose). The structure of WsaF bound to dTDP and dTDP-beta-l-rhamnose coupled to biochemical analysis identifies the residues that underlie catalysis and substrate recognition. We have constructed and tested by site-directed mutagenesis a model for acceptor recognition.
- Centre for Biomolecular Sciences, University of St. Andrews, North Haugh, St. Andrews, Fife KY16 9ST, UK.
Organizational Affiliation: