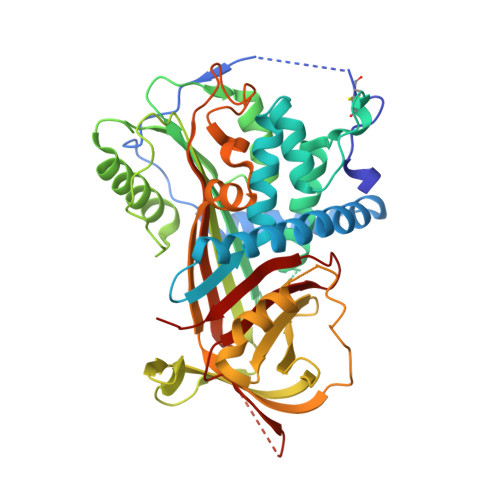
A Redox Switch in Angiotensinogen Modulates Angiotensin Release.
Zhou, A., Carrell, R.W., Murphy, M.P., Wei, Z., Yan, Y., Stanley, P.L., Stein, P.E., Pipkin, F.B., Read, R.J.(2010) Nature 468: 108
- PubMed: 20927107 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09505
- Primary Citation Related Structures:
2WXW, 2WXX, 2WXY, 2WXZ, 2WY0, 2WY1, 2X0B - PubMed Abstract:
Blood pressure is critically controlled by angiotensins, which are vasopressor peptides specifically released by the enzyme renin from the tail of angiotensinogen-a non-inhibitory member of the serpin family of protease inhibitors. Although angiotensinogen has long been regarded as a passive substrate, the crystal structures solved here to 2.1 Å resolution show that the angiotensin cleavage site is inaccessibly buried in its amino-terminal tail. The conformational rearrangement that makes this site accessible for proteolysis is revealed in our 4.4 Å structure of the complex of human angiotensinogen with renin. The co-ordinated changes involved are seen to be critically linked by a conserved but labile disulphide bridge. Here we show that the reduced unbridged form of angiotensinogen is present in the circulation in a near 40:60 ratio with the oxidized sulphydryl-bridged form, which preferentially interacts with receptor-bound renin. We propose that this redox-responsive transition of angiotensinogen to a form that will more effectively release angiotensin at a cellular level contributes to the modulation of blood pressure. Specifically, we demonstrate the oxidative switch of angiotensinogen to its more active sulphydryl-bridged form in the maternal circulation in pre-eclampsia-the hypertensive crisis of pregnancy that threatens the health and survival of both mother and child.
- Department of Haematology, Cambridge Institute for Medical Research, University of Cambridge, Hills Road, Cambridge CB2 0XY, UK. awz20@cam.ac.uk
Organizational Affiliation: