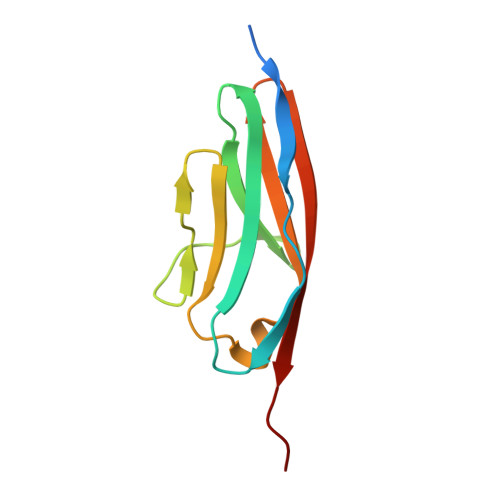
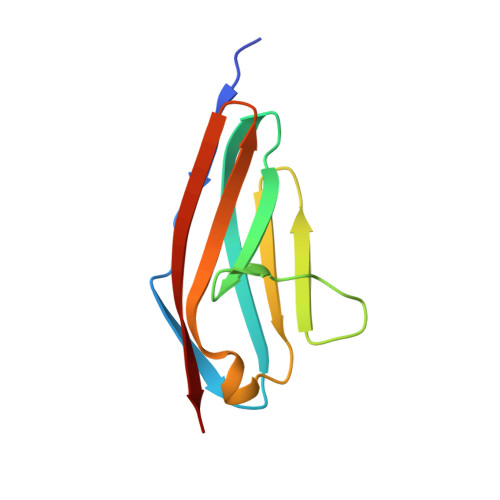
Structural Insight Into M-Band Assembly and Mechanics from the Titin-Obscurin-Like-1 Complex.
Pernigo, S., Fukuzawa, A., Bertz, M., Holt, M., Rief, M., Steiner, R.A., Gautel, M.(2010) Proc Natl Acad Sci U S A 107: 2908
- PubMed: 20133654 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0913736107
- Primary Citation Related Structures:
2WP3, 2WWK, 2WWM - PubMed Abstract:
In the sarcomeric M-band, the giant ruler proteins titin and obscurin, its small homologue obscurin-like-1 (obsl1), and the myosin cross-linking protein myomesin form a ternary complex that is crucial for the function of the M-band as a mechanical link. Mutations in the last titin immunoglobulin (Ig) domain M10, which interacts with the N-terminal Ig-domains of obscurin and obsl1, lead to hereditary muscle diseases. The M10 domain is unusual not only in that it is a frequent target of disease-linked mutations, but also in that it is the only currently known muscle Ig-domain that interacts with two ligands--obscurin and obsl1--in different sarcomeric subregions. Using x-ray crystallography, we show the structural basis for titin M10 interaction with obsl1 in a novel antiparallel Ig-Ig architecture and unravel the molecular basis of titin-M10 linked myopathies. The severity of these pathologies correlates with the disruption of the titin-obsl1/obscurin complex. Conserved signature residues at the interface account for differences in affinity that direct the cellular sorting in cardiomyocytes. By engineering the interface signature residues of obsl1 to obscurin, and vice versa, their affinity for titin can be modulated similar to the native proteins. In single-molecule force-spectroscopy experiments, both complexes yield at forces of around 30 pN, much lower than those observed for the mechanically stable Z-disk complex of titin and telethonin, suggesting why even moderate weakening of the obsl1/obscurin-titin links has severe consequences for normal muscle functions.
- King's College London BHF Research Excellence Centre, Randall Division for Cell and Molecular Biophysics and Cardiovascular Division, New Hunt's House, Guy's Campus, London SE1 1UL, United Kingdom.
Organizational Affiliation: