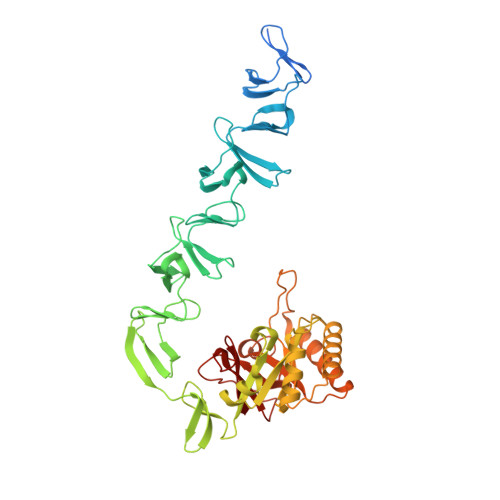
Insights Into Pneumococcal Fratricide from the Crystal Structures of the Modular Killing Factor Lytc.
Perez-Dorado, I., Gonzalez, A., Morales, M., Sanles, R., Striker, W., Vollmer, W., Mobashery, S., Garcia, J.L., Martinez-Ripoll, M., Garcia, P., Hermoso, J.A.(2010) Nat Struct Mol Biol 17: 576
- PubMed: 20400948 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1817
- Primary Citation Related Structures:
2WW5, 2WWC, 2WWD - PubMed Abstract:
The first structure of a pneumococcal autolysin, that of the LytC lysozyme, has been solved in ternary complex with choline and a pneumococcal peptidoglycan (PG) fragment. The active site of the hydrolase module is not fully exposed but is oriented toward the choline-binding module, which accounts for its unique in vivo features in PG hydrolysis, its activation and its regulatory mechanisms. Because of the unusual hook-shaped conformation of the multimodular protein, it is only able to hydrolyze non-cross-linked PG chains, an assertion validated by additional experiments. These results explain the activation of LytC by choline-binding protein D (CbpD) in fratricide, a competence-programmed mechanism of predation of noncompetent sister cells. The results provide the first structural insights to our knowledge into the critical and central function that LytC plays in pneumococcal virulence and explain a long-standing puzzle of how murein hydrolases can be controlled to avoid self-lysis during bacterial growth and division.
- Grupo de Cristalografía Macromolecular y Biología Estructural, Instituto Química-Física Rocasolano, Consejo Superior de Investigaciones Científicas, Madrid, Spain.
Organizational Affiliation: