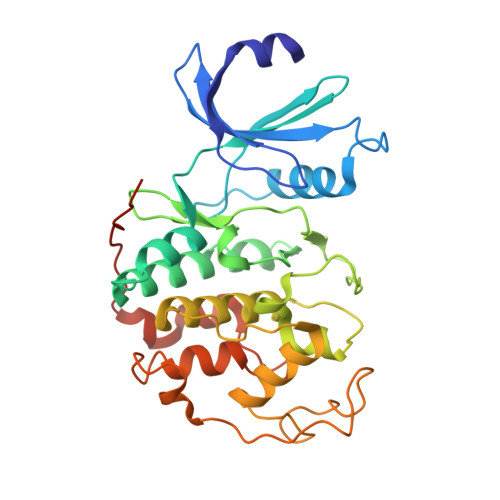
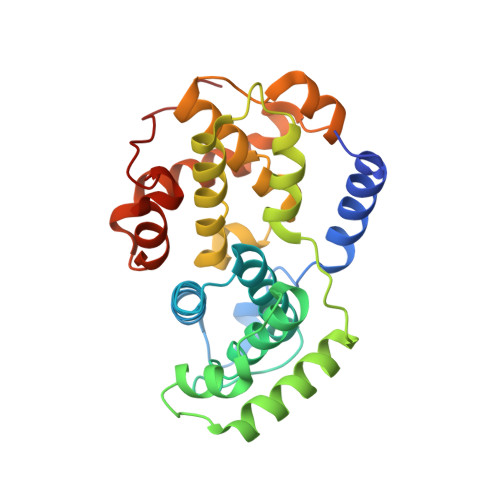
Optimization of 6,6-Dimethyl Pyrrolo[3,4-C]Pyrazoles: Identification of Pha-793887, a Potent Cdk Inhibitor Suitable for Intravenous Dosing.
Brasca, M.G., Albanese, C., Alzani, R., Amici, R., Avanzi, N., Ballinari, D., Bischoff, J., Borghi, D., Casale, E., Croci, V., Fiorentini, F., Isacchi, A., Mercurio, C., Nesi, M., Orsini, P., Pastori, W., Pesenti, E., Pevarello, P., Roussel, P., Varasi, M., Volpi, D., Vulpetti, A., Ciomei, M.(2010) Bioorg Med Chem 18: 1844
- PubMed: 20153204 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2010.01.042
- Primary Citation Related Structures:
2WPA - PubMed Abstract:
We have recently reported CDK inhibitors based on the 6-substituted pyrrolo[3,4-c]pyrazole core structure. Improvement of inhibitory potency against multiple CDKs, antiproliferative activity against cancer cell lines and optimization of the physico-chemical properties led to the identification of highly potent compounds. Compound 31 (PHA-793887) showed good efficacy in the human ovarian A2780, colon HCT-116 and pancreatic BX-PC3 carcinoma xenograft models and was well tolerated upon daily treatments by iv administration. It was identified as a drug candidate for clinical evaluation in patients with solid tumors.
- Nerviano Medical Sciences Srl, Business Unit Oncology, Viale Pasteur 10, 20014 Nerviano (MI), Italy. gabriella.brasca@nervianoms.com
Organizational Affiliation: