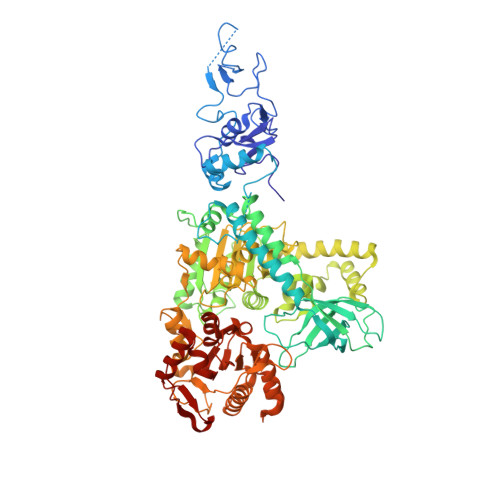
Unusual Bipartite Mode of Interaction between the Nonsense-Mediated Decay Factors, Upf1 and Upf2.
Clerici, M., Mourao, A., Gutsche, I., Gehring, N.H., Hentze, M.W., Kulozik, A., Kadlec, J., Sattler, M., Cusack, S.(2009) EMBO J 28: 2293
- PubMed: 19556969 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2009.175
- Primary Citation Related Structures:
2WJV, 2WJY - PubMed Abstract:
Nonsense-mediated decay (NMD) is a eukaryotic quality control mechanism that degrades mRNAs carrying premature stop codons. In mammalian cells, NMD is triggered when UPF2 bound to UPF3 on a downstream exon junction complex interacts with UPF1 bound to a stalled ribosome. We report structural studies on the interaction between the C-terminal region of UPF2 and intact UPF1. Crystal structures, confirmed by EM and SAXS, show that the UPF1 CH-domain is docked onto its helicase domain in a fixed configuration. The C-terminal region of UPF2 is natively unfolded but binds through separated alpha-helical and beta-hairpin elements to the UPF1 CH-domain. The alpha-helical region binds sixfold more weakly than the beta-hairpin, whereas the combined elements bind 80-fold more tightly. Cellular assays show that NMD is severely affected by mutations disrupting the beta-hairpin binding, but not by those only affecting alpha-helix binding. We propose that the bipartite mode of UPF2 binding to UPF1 brings the ribosome and the EJC in close proximity by forming a tight complex after an initial weak encounter with either element.
- European Molecular Biology Laboratory, Grenoble Outstation, Grenoble Cedex 9, France.
Organizational Affiliation: