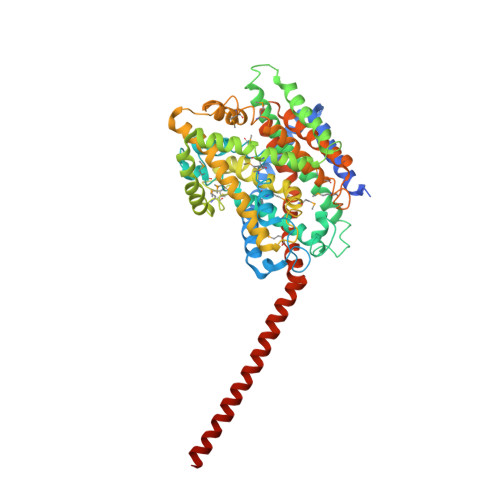
Molecular Basis of Transport and Regulation in the Na(+)/Betaine Symporter Betp.
Ressl, S., Terwisscha Van Scheltinga, A.C., Vonrhein, C., Ott, V., Ziegler, C.(2009) Nature 458: 47
- PubMed: 19262666 Search on PubMed
- DOI: https://doi.org/10.1038/nature07819
- Primary Citation Related Structures:
2WIT - PubMed Abstract:
Osmoregulated transporters sense intracellular osmotic pressure and respond to hyperosmotic stress by accumulation of osmolytes to restore normal hydration levels. Here we report the determination of the X-ray structure of a member of the family of betaine/choline/carnitine transporters, the Na(+)-coupled symporter BetP from Corynebacterium glutamicum, which is a highly effective osmoregulated uptake system for glycine betaine. Glycine betaine is bound in a tryptophan box occluded from both sides of the membrane with aromatic side chains lining the transport pathway. BetP has the same overall fold as three unrelated Na(+)-coupled symporters. Whereas these are crystallized in either the outward-facing or the inward-facing conformation, the BetP structure reveals a unique intermediate conformation in the Na(+)-coupled transport cycle. The trimeric architecture of BetP and the break in three-fold symmetry by the osmosensing C-terminal helices suggest a regulatory mechanism of Na(+)-coupled osmolyte transport to counteract osmotic stress.
- Max Planck Institute of Biophysics, Department of Structural Biology, 60438 Frankfurt am Main, Germany.
Organizational Affiliation: