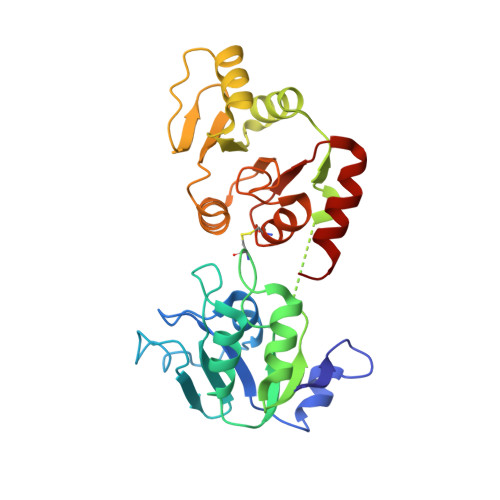
The Sars-Unique Domain (Sud) of Sars Coronavirus Contains Two Macrodomains that Bind G-Quadruplexes.
Tan, J., Vonrhein, C., Smart, O.S., Bricogne, G., Bollati, M., Kusov, Y., Hansen, G., Mesters, J.R., Schmidt, C.L., Hilgenfeld, R.(2009) PLoS Pathog 5: 428
- PubMed: 19436709 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1000428
- Primary Citation Related Structures:
2W2G, 2WCT - PubMed Abstract:
Since the outbreak of severe acute respiratory syndrome (SARS) in 2003, the three-dimensional structures of several of the replicase/transcriptase components of SARS coronavirus (SARS-CoV), the non-structural proteins (Nsps), have been determined. However, within the large Nsp3 (1922 amino-acid residues), the structure and function of the so-called SARS-unique domain (SUD) have remained elusive. SUD occurs only in SARS-CoV and the highly related viruses found in certain bats, but is absent from all other coronaviruses. Therefore, it has been speculated that it may be involved in the extreme pathogenicity of SARS-CoV, compared to other coronaviruses, most of which cause only mild infections in humans. In order to help elucidate the function of the SUD, we have determined crystal structures of fragment 389-652 ("SUD(core)") of Nsp3, which comprises 264 of the 338 residues of the domain. Both the monoclinic and triclinic crystal forms (2.2 and 2.8 A resolution, respectively) revealed that SUD(core) forms a homodimer. Each monomer consists of two subdomains, SUD-N and SUD-M, with a macrodomain fold similar to the SARS-CoV X-domain. However, in contrast to the latter, SUD fails to bind ADP-ribose, as determined by zone-interference gel electrophoresis. Instead, the entire SUD(core) as well as its individual subdomains interact with oligonucleotides known to form G-quadruplexes. This includes oligodeoxy- as well as oligoribonucleotides. Mutations of selected lysine residues on the surface of the SUD-N subdomain lead to reduction of G-quadruplex binding, whereas mutations in the SUD-M subdomain abolish it. As there is no evidence for Nsp3 entering the nucleus of the host cell, the SARS-CoV genomic RNA or host-cell mRNA containing long G-stretches may be targets of SUD. The SARS-CoV genome is devoid of G-stretches longer than 5-6 nucleotides, but more extended G-stretches are found in the 3'-nontranslated regions of mRNAs coding for certain host-cell proteins involved in apoptosis or signal transduction, and have been shown to bind to SUD in vitro. Therefore, SUD may be involved in controlling the host cell's response to the viral infection. Possible interference with poly(ADP-ribose) polymerase-like domains is also discussed.
- Institute of Biochemistry, Center for Structural and Cell Biology in Medicine, University of Lübeck, Lübeck, Germany.
Organizational Affiliation: