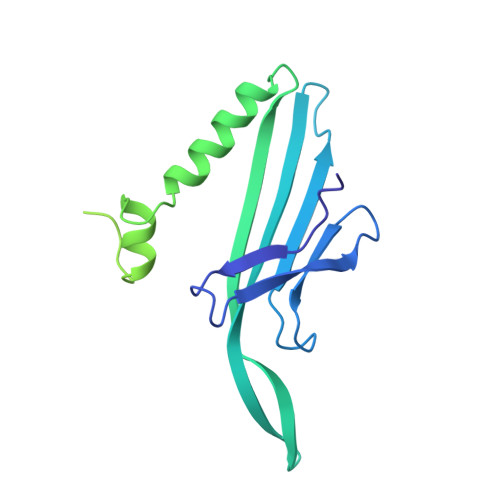
Structure and Stability of Icosahedral Particles of a Covalent Coat Protein Dimer of Bacteriophage MS2.
Plevka, P., Tars, K., Liljas, L.(2009) Protein Sci 18: 1653
- PubMed: 19521994 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.184
- Primary Citation Related Structures:
2WBH - PubMed Abstract:
Particles formed by the bacteriophage MS2 coat protein mutants with insertions in their surface loops induce a strong immune response against the inserted epitopes. The covalent dimers created by fusion of two copies of the coat protein gene are more tolerant to various insertions into the surface loops than the single subunits. We determined a 4.7-A resolution crystal structure of an icosahedral particle assembled from covalent dimers and compared its stability with wild-type virions. The structure resembled the wild-type virion except for the intersubunit linker regions. The covalent dimer orientation was random with respect to both icosahedral twofold and quasi-twofold symmetry axes. A fraction of the particles was unstable in phosphate buffer because of assembly defects. Our results provide a structural background for design of modified covalent coat protein dimer subunits for use in immunization.
- Department of Cell and Molecular Biology, Uppsala University, SE-751 24 Uppsala, Sweden. pp2cz@yahoo.com
Organizational Affiliation: