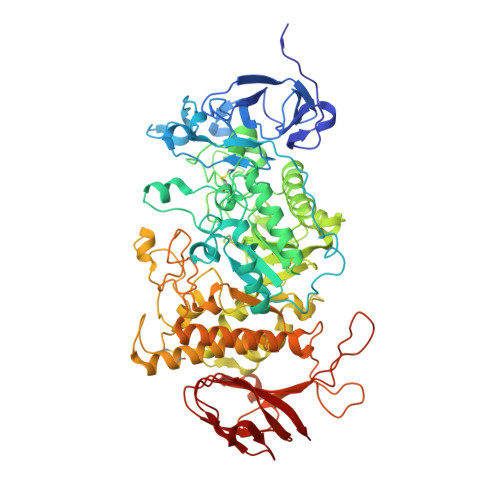
Structural Insight Into the Bifunctional Mechanism of the Glycogen-Debranching Enzyme Trex from the Archaeon Sulfolobus Solfataricus.
Woo, E., Lee, S., Cha, H., Park, J., Yoon, S., Song, H., Park, K.(2008) J Biological Chem 283: 28641
- PubMed: 18703518
- DOI: https://doi.org/10.1074/jbc.M802560200
- Primary Citation Related Structures:
2VNC, 2VR5, 2VUY - PubMed Abstract:
TreX is an archaeal glycogen-debranching enzyme that exists in two oligomeric states in solution, as a dimer and tetramer. Unlike its homologs, TreX from Sulfolobus solfataricus shows dual activities for alpha-1,4-transferase and alpha-1,6-glucosidase. To understand this bifunctional mechanism, we determined the crystal structure of TreX in complex with an acarbose ligand. The acarbose intermediate was covalently bound to Asp363, occupying subsites -1 to -3. Although generally similar to the monomeric structure of isoamylase, TreX exhibits two different active-site configurations depending on its oligomeric state. The N terminus of one subunit is located at the active site of the other molecule, resulting in a reshaping of the active site in the tetramer. This is accompanied by a large shift in the "flexible loop" (amino acids 399-416), creating connected holes inside the tetramer. Mutations in the N-terminal region result in a sharp increase in alpha-1,4-transferase activity and a reduced level of alpha-1,6-glucosidase activity. On the basis of geometrical analysis of the active site and mutational study, we suggest that the structural lid (acids 99-97) at the active site generated by the tetramerization is closely associated with the bifunctionality and in particular with the alpha-1,4-transferase activity. These results provide a structural basis for the modulation of activities upon TreX oligomerization that may represent a common mode of action for other glycogen-debranching enzymes in higher organisms.
- Translational Research Center, Korea Research Institute of Bioscience and Biotechnology, 111 Gwahangno, Yuseong-gu, Daejeon 305-806.
Organizational Affiliation: