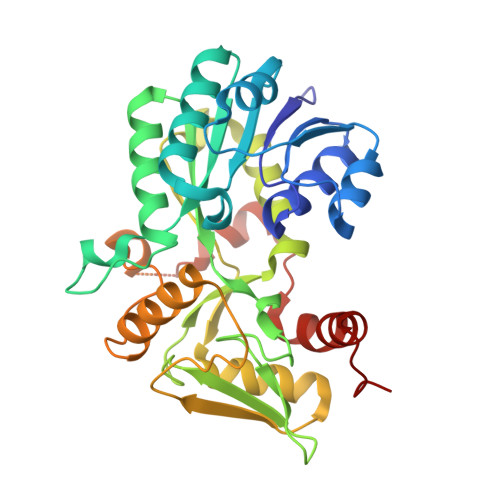
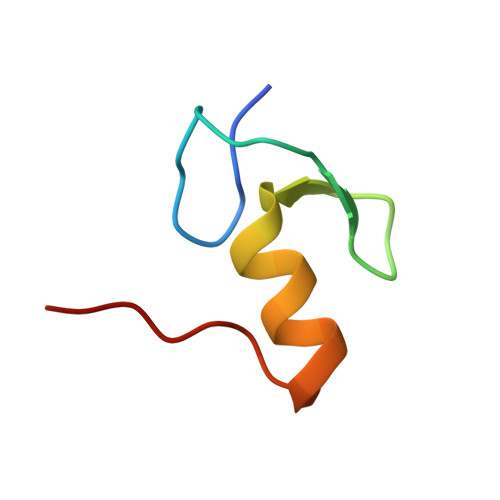
Structural Analysis of the Recognition of the Negative Regulator Nmra and DNA by the Zinc Finger from the Gata-Type Transcription Factor Area.
Kotaka, M., Johnson, C., Lamb, H.K., Hawkins, A.R., Ren, J., Stammers, D.K.(2008) J Mol Biology 381: 373
- PubMed: 18602114 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.05.077
- Primary Citation Related Structures:
2VUS, 2VUT, 2VUU - PubMed Abstract:
Amongst the most common protein motifs in eukaryotes are zinc fingers (ZFs), which, although largely known as DNA binding modules, also can have additional important regulatory roles in forming protein:protein interactions. AreA is a transcriptional activator central to nitrogen metabolism in Aspergillus nidulans. AreA contains a GATA-type ZF that has a competing dual recognition function, binding either DNA or the negative regulator NmrA. We report the crystal structures of three AreA ZF-NmrA complexes including two with bound NAD(+) or NADP(+). The molecular recognition of AreA ZF-NmrA involves binding of the ZF to NmrA via hydrophobic and hydrogen bonding interactions through helices alpha1, alpha6 and alpha11. Comparison with an earlier NMR solution structure of AreA ZF-DNA complex by overlap of the AreA ZFs shows that parts of helices alpha6 and alpha11 of NmrA are positioned close to the GATA motif of the DNA, mimicking the major groove of DNA. The extensive overlap of DNA with NmrA explains their mutually exclusive binding to the AreA ZF. The presence of bound NAD(+)/NADP(+) in the NmrA-AreaA ZF complex, however, causes minimal structural changes. Thus, any regulatory effects on AreA function mediated by the binding of oxidised nicotinamide dinucleotides to NmrA in the NmrA-AreA ZF complex appear not to be modulated via protein conformational rearrangements.
- Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Oxford, OX3 7BN, UK.
Organizational Affiliation: