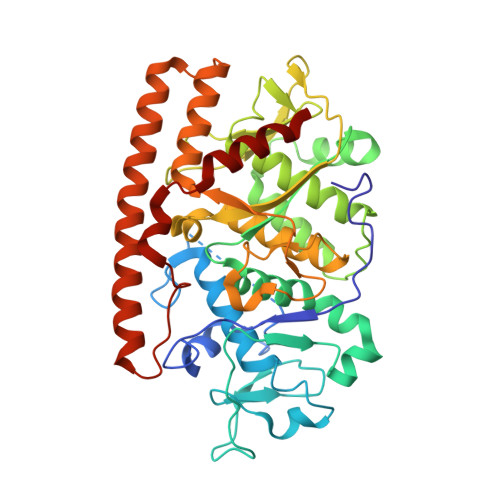
Crystal structure of the avian reovirus inner capsid protein sigmaA.
Guardado-Calvo, P., Vazquez-Iglesias, L., Martinez-Costas, J., Llamas-Saiz, A.L., Schoehn, G., Fox, G.C., Hermo-Parrado, X.L., Benavente, J., van Raaij, M.J.(2008) J Virol 82: 11208-11216
- PubMed: 18799570 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00733-08
- Primary Citation Related Structures:
2VAK - PubMed Abstract:
Avian reovirus, an important avian pathogen, expresses eight structural and four nonstructural proteins. The structural sigmaA protein is a major component of the inner capsid, clamping together lambdaA building blocks. sigmaA has also been implicated in the resistance of avian reovirus to the antiviral action of interferon by strongly binding double-stranded RNA in the host cell cytoplasm and thus inhibiting activation of the double-stranded RNA-dependent protein kinase. We have solved the structure of bacterially expressed sigmaA by molecular replacement and refined it using data to 2.3-A resolution. Twelve sigmaA molecules are present in the P1 unit cell, arranged as two short double helical hexamers. A positively charged patch is apparent on the surface of sigmaA on the inside of this helix and mutation of either of two key arginine residues (Arg155 and Arg273) within this patch abolishes double-stranded RNA binding. The structural data, together with gel shift assay, electron microscopy, and sedimentation velocity centrifugation results, provide evidence for cooperative binding of sigmaA to double-stranded RNA. The minimal length of double-stranded RNA required for sigmaA binding was observed to be 14 to 18 bp.
- Departamento de Bioquímica e Bioloxía Molecular, Facultade de Farmacia, Universidade de Santiago de Compostela, E-15782 Santiago de Compostela, Spain.
Organizational Affiliation: