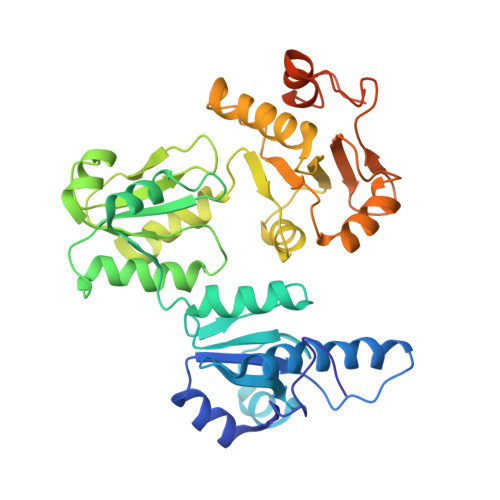
Characterization of Human Cardiac Calsequestrin and its Deleterious Mutants.
Kim, E., Youn, B., Kemper, L., Campbell, C., Milting, H., Varsanyi, M., Kang, C.(2007) J Mol Biology 373: 1047
- PubMed: 17881003 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.08.055
- Primary Citation Related Structures:
2VAF - PubMed Abstract:
Mutations of conserved residues of human cardiac calsequestrin (hCSQ2), a high-capacity, low-affinity Ca2+-binding protein in the sarcoplasmic reticulum, have been associated with catecholamine-induced polymorphic ventricular tachycardia (CPVT). In order to understand the molecular mechanism and pathophysiological link between these CPVT-related missense mutations of hCSQ2 and the resulting arrhythmias, we generated three CPVT-causing mutants of hCSQ2 (R33Q, L167H, and D307H) and two non-pathological mutants (T66A and V76M) and investigated the effect of these mutations. In addition, we determined the crystal structure of the corresponding wild-type hCSQ2 to gain insight into the structural effects of those mutations. Our data show clearly that all three CPVT-related mutations lead to significant reduction in Ca2+-binding capacity in spite of the similarity of their secondary structures to that of the wild-type hCSQ2. Light-scattering experiments indicate that the Ca2+-dependent monomer-polymer transitions of the mutants are quite different, confirming that the linear polymerization behavior of CSQ is linked directly to its high-capacity Ca2+ binding. R33Q and D307H mutations result in a monomer that appears to be unable to form a properly oriented dimer. On the other hand, the L167H mutant has a disrupted hydrophobic core in domain II, resulting in high molecular aggregates, which cannot respond to Ca2+. Although one of the non-pathological mutants, T66A, shares characteristics with the wild-type, the other null mutant, V76M, shows significantly altered Ca2+-binding and polymerization behaviors, calling for careful reconsideration of its status.
- School of Molecular Biosciences, Washington State University Pullman, WA 99164-4660, USA.
Organizational Affiliation: