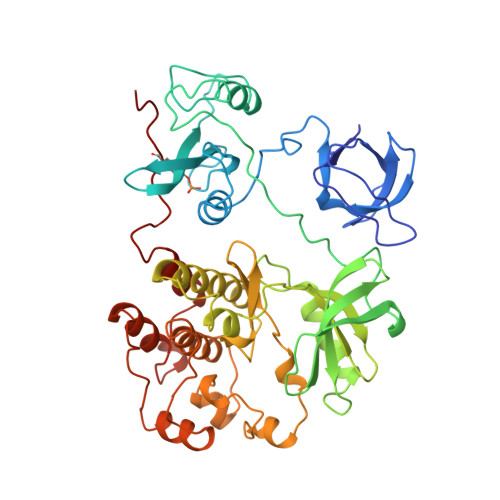
Crystal structures of c-Src reveal features of its autoinhibitory mechanism.
Xu, W., Doshi, A., Lei, M., Eck, M.J., Harrison, S.C.(1999) Mol Cell 3: 629-638
- PubMed: 10360179 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(00)80356-1
- Primary Citation Related Structures:
2SRC - PubMed Abstract:
Src family kinases are maintained in an assembled, inactive conformation by intramolecular interactions of their SH2 and SH3 domains. Full catalytic activity requires release of these restraints as well as phosphorylation of Tyr-416 in the activation loop. In previous structures of inactive Src kinases, Tyr-416 and flanking residues are disordered. We report here four additional c-Src structures in which this segment adopts an ordered but inhibitory conformation. The ordered activation loop forms an alpha helix that stabilizes the inactive conformation of the kinase domain, blocks the peptide substrate-binding site, and prevents Tyr-416 phosphorylation. Disassembly of the regulatory domains, induced by SH2 or SH3 ligands, or by dephosphorylation of Tyr-527, could lead to exposure and phosphorylation of Tyr-416.
- Laboratory of Molecular Medicine, Children's Hospital, Boston, Massachusetts 02115, USA.
Organizational Affiliation: