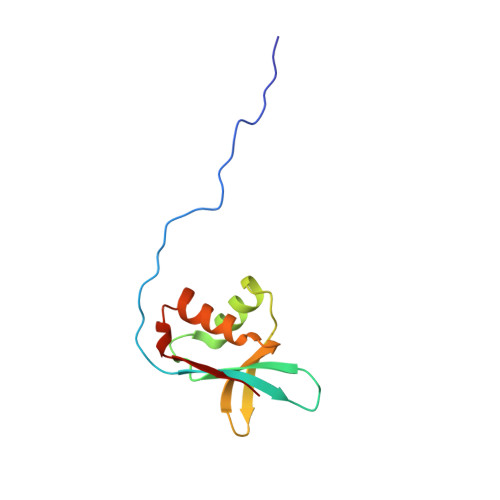
Molecular Landscape of the Ribosome Pre-initiation Complex during mRNA Scanning: Structural Role for eIF3c and Its Control by eIF5
Obayashi, E., Luna, R.E., Nagata, T., Martin-Marcos, P., Hiraishi, H., Singh, C.R., Erzberger, J.P., Zhang, F., Arthanari, H., Morris, J., Pellarin, R., Moore, C., Harmon, I., Papadopoulos, E., Yoshida, H., Nasr, M.L., Unzai, S., Thompson, B., Aube, E., Hustak, S., Stengel, F., Dagraca, E., Ananbandam, A., Gao, P., Urano, T., Hinnebusch, A.G., Wagner, G., Asano, K.(2017) Cell Rep 18: 2651-2663
- PubMed: 28297669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2017.02.052
- Primary Citation Related Structures:
2RVH, 5H7U - PubMed Abstract:
During eukaryotic translation initiation, eIF3 binds the solvent-accessible side of the 40S ribosome and recruits the gate-keeper protein eIF1 and eIF5 to the decoding center. This is largely mediated by the N-terminal domain (NTD) of eIF3c, which can be divided into three parts: 3c0, 3c1, and 3c2. The N-terminal part, 3c0, binds eIF5 strongly but only weakly to the ribosome-binding surface of eIF1, whereas 3c1 and 3c2 form a stoichiometric complex with eIF1. 3c1 contacts eIF1 through Arg-53 and Leu-96, while 3c2 faces 40S protein uS15/S13, to anchor eIF1 to the scanning pre-initiation complex (PIC). We propose that the 3c0:eIF1 interaction diminishes eIF1 binding to the 40S, whereas 3c0:eIF5 interaction stabilizes the scanning PIC by precluding this inhibitory interaction. Upon start codon recognition, interactions involving eIF5, and ultimately 3c0:eIF1 association, facilitate eIF1 release. Our results reveal intricate molecular interactions within the PIC, programmed for rapid scanning-arrest at the start codon.
- Shimane University School of Medicine, Izumo, Shimane 690-8504, Japan.
Organizational Affiliation: