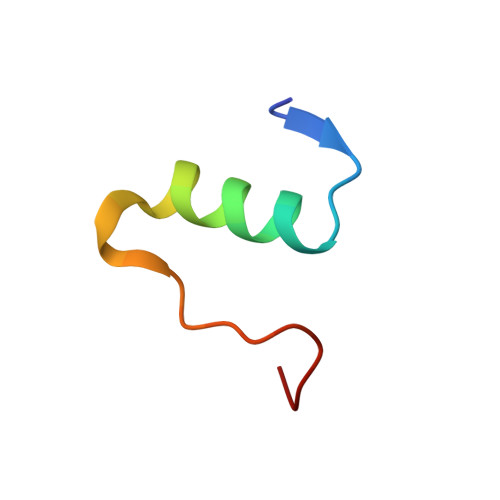
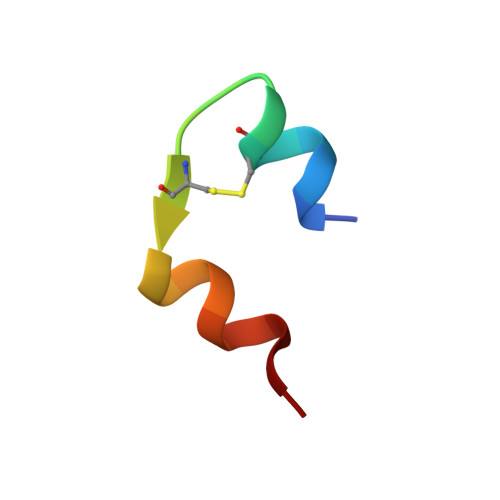NMR structure of biosynthetic engineered human insulin monomer B31(Lys)-B32(Arg) in water/acetonitrile solution. Comparison with the solution structure of native human insulin monomer
Bocian, W., Borowicz, P., Sitkowski, J., Tarnowska, A., Bednarek, E., Bogiel, M., Kozerski, L.(2008) Biopolymers 89: 820-830
- PubMed: 18491415 Search on PubMed
- DOI: https://doi.org/10.1002/bip.21018
- Primary Citation Related Structures:
2RN5 - PubMed Abstract:
A solution NMR-derived structure of a new long -acting, B31(Lys)-B32(Arg) (LysArg), engineered human insulin monomer, in H(2)O/CD(3)CN, 65/35 vol %, pH 3.6, is presented and compared with the available X-ray structure of a monomer that forms part of a hexamer (Smith, et al., Acta Crystallogr D 2003, 59, 474) and with NMR structure of human insulin in the same solvent (Bocian, et al., J Biomol NMR 2008, 40, 55-64). Detailed analysis using PFGSE NMR (Pulsed Field Gradient Spin Echo NMR) in dilution experiments and CSI analysis prove that the structure is monomeric in the concentration range 0.1-3 mM. The presence of long-range interstrand NOEs in a studied structure, relevant to the distances found in the crystal structure of the monomer, provides the evidence for conservation of the tertiary structure. Therefore the results suggest that this solvent system is a suitable medium for studying the native conformation of the protein, especially in situations (as found for insulins) in which extensive aggregation renders structure elucidations in water difficult or impossible. Starting from the structures calculated by the program CYANA, two different molecular dynamics (MD) simulated annealing refinement protocols were applied, either using the program AMBER in vacuum (AMBER_VC), or including a generalized Born solvent model (AMBER_GB). Here we present another independent evidence to the one presented recently by us (Bocian et al., J Biomol NMR 2008, 40, 55-64), that in water/acetonitrile solvent detailed structural and dynamic information can be obtained for important proteins that are naturally present as oligomers under native conditions.
- National Medicines Institute, 00-725 Warsaw, Poland.
Organizational Affiliation: