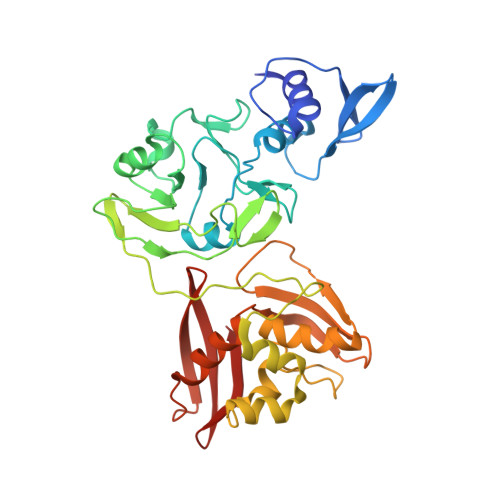
Structural and functional analyses of the severe acute respiratory syndrome coronavirus endoribonuclease Nsp15.
Bhardwaj, K., Palaninathan, S., Alcantara, J.M., Yi, L.L., Guarino, L., Sacchettini, J.C., Kao, C.C.(2008) J Biological Chem 283: 3655-3664
- PubMed: 18045871 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M708375200
- Primary Citation Related Structures:
2RHB - PubMed Abstract:
The severe acute respiratory syndrome (SARS) coronavirus encodes several RNA-processing enzymes that are unusual for RNA viruses, including Nsp15 (nonstructural protein 15), a hexameric endoribonuclease that preferentially cleaves 3' of uridines. We solved the structure of a catalytically inactive mutant version of Nsp15, which was crystallized as a hexamer. The structure contains unreported flexibility in the active site of each subunit. Substitutions in the active site residues serine 293 and proline 343 allowed Nsp15 to cleave at cytidylate, whereas mutation of leucine 345 rendered Nsp15 able to cleave at purines as well as pyrimidines. Mutations that targeted the residues involved in subunit interactions generally resulted in the formation of catalytically inactive monomers. The RNA-binding residues were mapped by a method linking reversible cross-linking, RNA affinity purification, and peptide fingerprinting. Alanine substitution of several residues in the RNA-contacting portion of Nsp15 did not affect hexamer formation but decreased the affinity of RNA binding and reduced endonuclease activity. This suggests a model for Nsp15 hexamer interaction with RNA.
- Department of Biochemistry and Biophysics, Texas A & M University, College Station, Texas 77843-2128.
Organizational Affiliation: