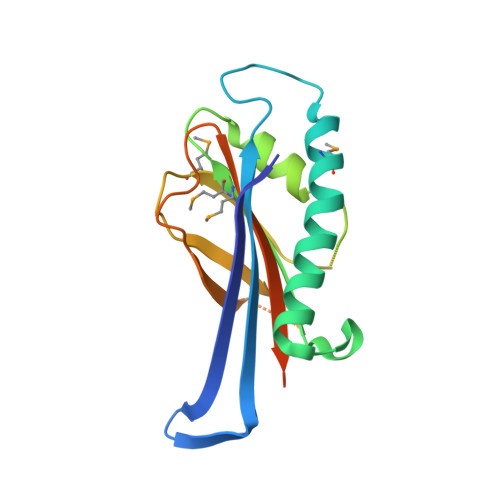
Structures of the first representatives of Pfam family PF06684 (DUF1185) reveal a novel variant of the Bacillus chorismate mutase fold and suggest a role in amino-acid metabolism.
Bakolitsa, C., Kumar, A., Jin, K.K., McMullan, D., Krishna, S.S., Miller, M.D., Abdubek, P., Acosta, C., Astakhova, T., Axelrod, H.L., Burra, P., Carlton, D., Chen, C., Chiu, H.J., Clayton, T., Das, D., Deller, M.C., Duan, L., Elias, Y., Ellrott, K., Ernst, D., Farr, C.L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Grzechnik, S.K., Han, G.W., Jaroszewski, L., Johnson, H.A., Klock, H.E., Knuth, M.W., Kozbial, P., Marciano, D., Morse, A.T., Murphy, K.D., Nigoghossian, E., Nopakun, A., Okach, L., Paulsen, J., Puckett, C., Reyes, R., Rife, C.L., Sefcovic, N., Tien, H.J., Trame, C.B., Trout, C.V., van den Bedem, H., Weekes, D., White, A., Xu, Q., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1182-1189
- PubMed: 20944209 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109050647
- Primary Citation Related Structures:
2QTP, 3BYQ - PubMed Abstract:
The crystal structures of BB2672 and SPO0826 were determined to resolutions of 1.7 and 2.1 Å by single-wavelength anomalous dispersion and multiple-wavelength anomalous dispersion, respectively, using the semi-automated high-throughput pipeline of the Joint Center for Structural Genomics (JCSG) as part of the NIGMS Protein Structure Initiative (PSI). These proteins are the first structural representatives of the PF06684 (DUF1185) Pfam family. Structural analysis revealed that both structures adopt a variant of the Bacillus chorismate mutase fold (BCM). The biological unit of both proteins is a hexamer and analysis of homologs indicates that the oligomer interface residues are highly conserved. The conformation of the critical regions for oligomerization appears to be dependent on pH or salt concentration, suggesting that this protein might be subject to environmental regulation. Structural similarities to BCM and genome-context analysis suggest a function in amino-acid synthesis.
- Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.
Organizational Affiliation: