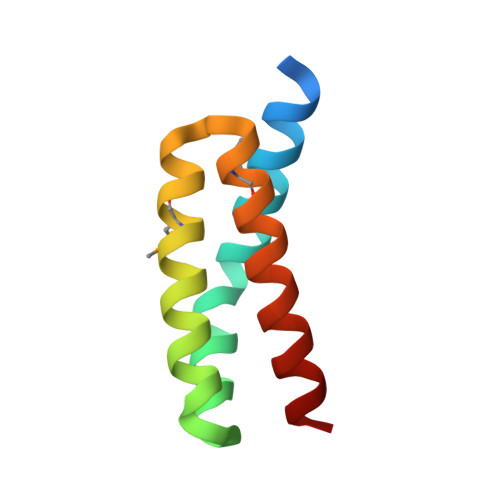
Staphylococcal complement inhibitor: structure and active sites.
Rooijakkers, S.H., Milder, F.J., Bardoel, B.W., Ruyken, M., van Strijp, J.A., Gros, P.(2007) J Immunol 179: 2989-2998
- PubMed: 17709514 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.179.5.2989
- Primary Citation Related Structures:
2QFF - PubMed Abstract:
The pathogenic bacterium Staphylococcus aureus counteracts the host immune defense by excretion of the 85 residue staphylococcal complement inhibitor (SCIN). SCIN inhibits the central complement convertases; thereby, it reduces phagocytosis following opsonization and efficiently blocks all downstream effector functions. In this study, we present the crystal structure of SCIN at 1.8 A resolution and the identification of its active site. Functional characterization of structure based chimeric proteins, consisting of SCIN and the structurally but nonfunctional homologue open reading frame-D, indicate an 18-residue segment (Leu-31-Gly-48) crucial for SCIN activity. In all complement activation pathways, chimeras lacking these SCIN residues completely fail to inhibit production of the potent mediator of inflammation C5a. Inhibition of alternative pathway-mediated opsonization (C3b deposition) and formation of the lytic membrane attack complex (C5b-9 deposition) are strongly reduced for these chimeras as well. For inhibition of the classical/lectin pathway-mediated C3b and C5b-9 deposition, the same residues are critical although additional sites are involved. These chimeras also display reduced capacity to stabilize the C3 convertases of both the alternative and the classical/lectin pathway indicating the stabilizing effect is pivotal for the complement inhibitory activity of SCIN. Because SCIN specifically and efficiently inhibits complement, it has a high potential in anti-inflammatory therapy. Our data are a first step toward the development of a second generation molecule suitable for such therapeutic complement intervention.
- Medical Microbiology, University Medical Center Utrecht, Heidelberglaan 100, Utrecht, The Netherlands. s.h.m.rooijakkers@umcutrecht.nl
Organizational Affiliation: