Design of protein function leaps by directed domain interface evolution.
Huang, J., Koide, A., Makabe, K., Koide, S.(2008) Proc Natl Acad Sci U S A 105: 6578-6583
- PubMed: 18445649 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0801097105
- Primary Citation Related Structures:
2QBW - PubMed Abstract:
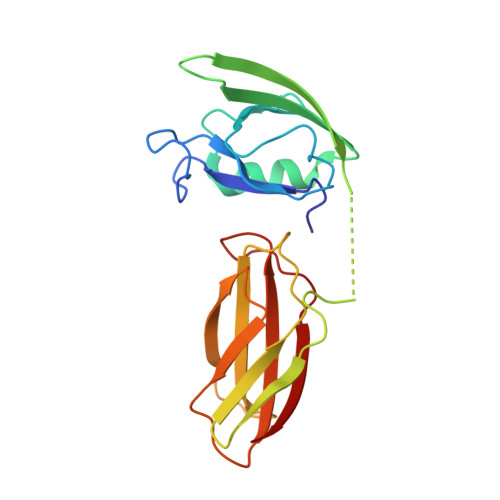
Most natural proteins performing sophisticated tasks contain multiple domains where an active site is located at the domain interface. Comparative structural analyses suggest that major leaps in protein function occur through gene recombination events that connect two or more protein domains to generate a new active site, frequently occurring at the newly created domain interface. However, such functional leaps by combination of unrelated domains have not been directly demonstrated. Here we show that highly specific and complex protein functions can be generated by joining a low-affinity peptide-binding domain with a functionally inert second domain and subsequently optimizing the domain interface. These directed evolution processes dramatically enhanced both affinity and specificity to a level unattainable with a single domain, corresponding to >500-fold and >2,000-fold increases of affinity and specificity, respectively. An x-ray crystal structure revealed that the resulting "affinity clamp" had clamshell architecture as designed, with large additional binding surface contributed by the second domain. The affinity clamps having a single-nanomolar dissociation constant outperformed a monoclonal antibody in immunochemical applications. This work establishes evolutionary paths from isolated domains with primitive function to multidomain proteins with sophisticated function and introduces a new protein-engineering concept that allows for the generation of highly functional affinity reagents to a predefined target. The prevalence and variety of natural interaction domains suggest that numerous new functions can be designed by using directed domain interface evolution.
- Department of Biochemistry and Molecular Biology, University of Chicago, 929 East 57th Street, Chicago, IL 60637, USA.
Organizational Affiliation: