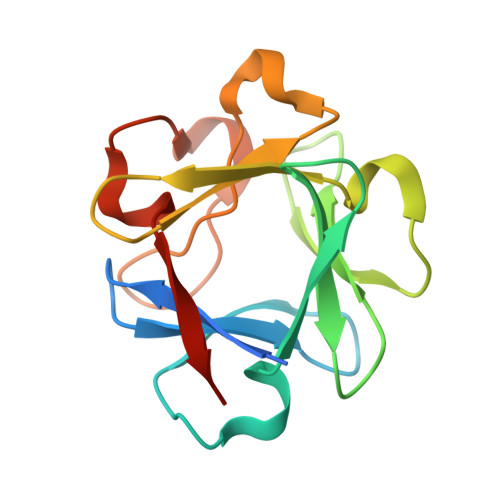
Structure of a highly stable mutant of human fibroblast growth factor 1.
Szlachcic, A., Zakrzewska, M., Krowarsch, D., Os, V., Helland, R., Smalas, A.O., Otlewski, J.(2009) Acta Crystallogr D Biol Crystallogr 65: 67-73
- PubMed: 19153468 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444908039486
- Primary Citation Related Structures:
2Q9X - PubMed Abstract:
Fibroblast growth factors (FGFs) are involved in diverse cellular processes such as cell migration, angiogenesis, osteogenesis, wound healing and embryonic and foetal development. Human acidic fibroblast growth factor (FGF-1) is the only member of the FGF family that binds with high affinity to all four FGF receptors and thus is considered to be the human mitogen with the broadest specificity. However, pharmacological applications of FGF-1 are limited owing to its low stability. It has previously been reported that the introduction of single mutations can significantly improve the stability of FGF-1 and its resistance to proteolytic degradation. Here, the structure of the Q40P/S47I/H93G triple mutant of FGF-1, which exhibits much higher stability, a prolonged half-life and enhanced mitogenic activity, is presented. Compared with the wild-type structure, three localized conformational changes in the stable triple mutant were observed, which is in agreement with the perfect energetic additivity of the single mutations described in a previous study. The huge change in FGF-1 stability (the denaturation temperature increased by 21.5 K, equivalent to DeltaDeltaG(den) = 24.3 kJ mol(-1)) seems to result from the formation of a short 3(10)-helix (position 40), an improvement in the propensity of amino acids to form beta-sheets (position 47) and the rearrangement of a local hydrogen-bond network (positions 47 and 93).
- Department of Protein Engineering, Faculty of Biotechnology, University of Wroclaw, Wroclaw, Poland.
Organizational Affiliation: