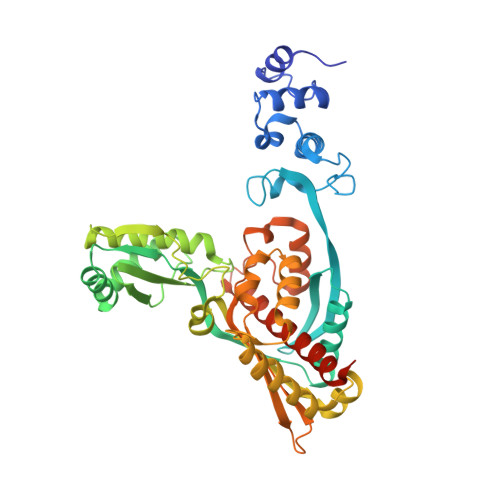
Design and synthesis of novel, conformationally restricted HMG-CoA reductase inhibitors.
Pfefferkorn, J.A., Choi, C., Song, Y., Trivedi, B.K., Larsen, S.D., Askew, V., Dillon, L., Hanselman, J.C., Lin, Z., Lu, G., Robertson, A., Sekerke, C., Auerbach, B., Pavlovsky, A., Harris, M.S., Bainbridge, G., Caspers, N.(2007) Bioorg Med Chem Lett 17: 4531-4537
- PubMed: 17574411 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.05.097
- Primary Citation Related Structures:
2Q6B, 2Q6C - PubMed Abstract:
Using structure-based design, a novel series of conformationally restricted, pyrrole-based inhibitors of HMG-CoA reductase were discovered. Leading analogs demonstrated potent inhibition of cholesterol synthesis in both in vitro and in vivo models and may be useful for the treatment of hypercholesterolemia and related lipid disorders.
- Pfizer Global Research & Development, Michigan Laboratories, 2800 Plymouth Road, Ann Arbor, MI 48105, USA. jeffrey.a.pfefferkorn@pfizer.com
Organizational Affiliation: