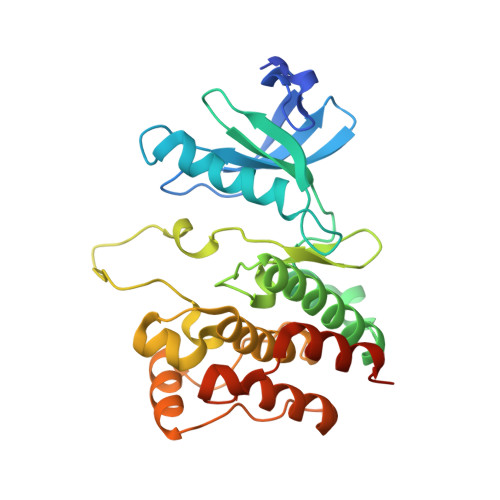
Classifying protein kinase structures guides use of ligand-selectivity profiles to predict inactive conformations: Structure of lck/imatinib complex.
Jacobs, M.D., Caron, P.R., Hare, B.J.(2007) Proteins 70: 1451-1460
- PubMed: 17910071 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21633
- Primary Citation Related Structures:
2PL0 - PubMed Abstract:
We report a clustering of public human protein kinase structures based on the conformations of two structural elements, the activation segment and the C-helix, revealing three discrete clusters. One cluster includes kinases in catalytically active conformations. Each of the other clusters contains a distinct inactive conformation. Typically, kinases adopt at most one of the inactive conformations in available X-ray structures, implying that one of the conformations is preferred for many kinases. The classification is consistent with selectivity profiles of several well-characterized kinase inhibitors. We show further that inhibitor selectivity profiles guide kinase classification. For example, selective inhibition of lck among src-family kinases by imatinib (Gleevec) suggests that the relative stabilities of inactive conformations of lck are different from other src-family kinases. We report the X-ray structure of the lck/imatinib complex, confirming that the conformation adopted by lck is distinct from other structurally-characterized src-family kinases and instead resembles kinases abl1 and kit in complex with imatinib. Our classification creates new paths for designing small-molecule inhibitors.
- Vertex Pharmaceuticals Incorporated, 130 Waverly Street, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: