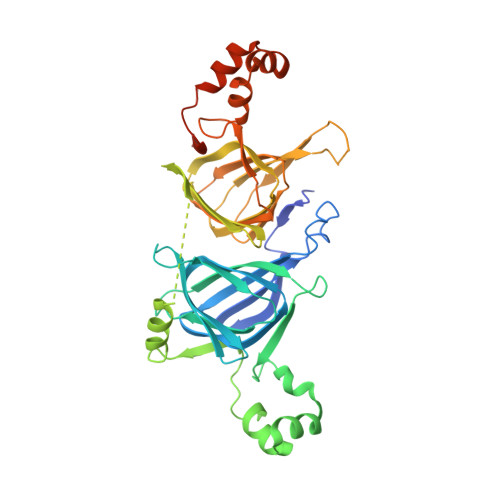
Structure of phaseolin at 2.2 A resolution. Implications for a common vicilin/legumin structure and the genetic engineering of seed storage proteins.
Lawrence, M.C., Izard, T., Beuchat, M., Blagrove, R.J., Colman, P.M.(1994) J Mol Biology 238: 748-776
- PubMed: 8182747 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.1333
- Primary Citation Related Structures:
2PHL - PubMed Abstract:
The refinement to 2.2 A resolution of the three-dimensional structure of the seed storage protein phaseolin from the French bean (Phaseolus vulgaris) via an alternative crystal form is described. The refined structure reveals details of the molecule hitherto unobserved and in particular we identify the structural role of conserved residues within the broader 7 S (vicilin) family of seed storage proteins. On this basis we are able to postulate a canonical model for the structure of the 7 S proteins. This model in turn provides a means for interpreting the structure of the 11 S (legumin) family of seed storage proteins, for which no X-ray diffraction data are available. The 11 S proteins are shown to bear a much closer relationship to the 7 S proteins than was previously recognized. The canonical model of the 7 S protein structure also provides a basis for proposing engineered mutations of these proteins with the goal of enhancing nutritional and functional properties.
- Biomolecular Research Institute, Parkville, Victoria, Australia.
Organizational Affiliation: