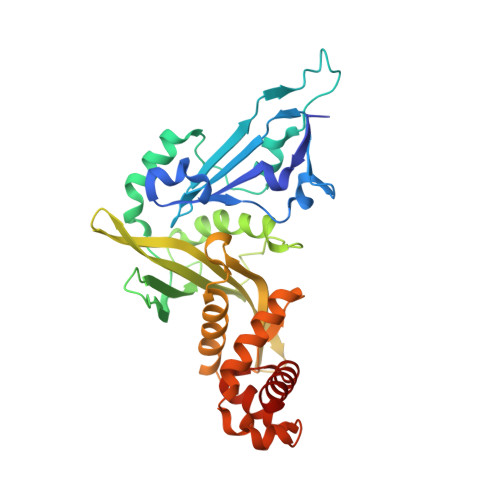
The crystal structure of human histone acetyltransferase 1 (HAT1) in complex with acetylcoenzyme A and histone peptide H4
Wu, H., Min, J., Zeng, H., Loppnau, P., Weigelt, J., Sundstrom, M., Arrowsmith, C.H., Edwards, A.M., Bochkarev, A., Plotnikov, A.N.To be published.