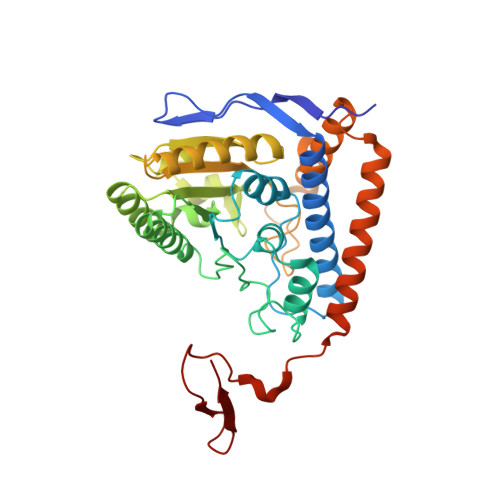
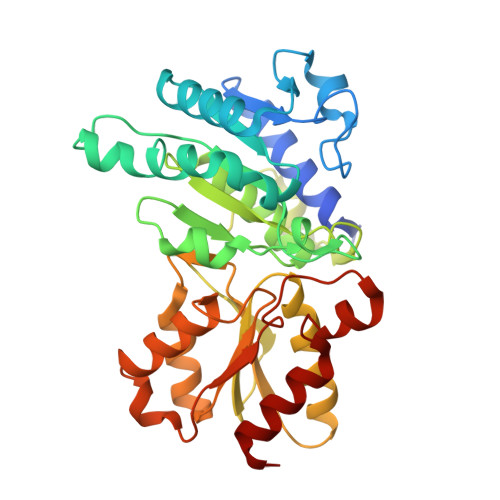
Phosphorylation of Serine 264 Impedes Active Site Accessibility in the E1 Component of the Human Pyruvate Dehydrogenase Multienzyme Complex
Seifert, F., Ciszak, E.M., Korotchkina, L.G., Golbik, R., Spinka, M., Dominiak, P.M., Sidhu, S., Brauer, J., Patel, M.S., Tittmann, K.(2007) Biochemistry 46: 6277-6287
- PubMed: 17474719 Search on PubMed
- DOI: https://doi.org/10.1021/bi700083z
- Primary Citation Related Structures:
2OZL - PubMed Abstract:
At the junction of glycolysis and the Krebs cycle in cellular metabolism, the pyruvate dehydrogenase multienzyme complex (PDHc) catalyzes the oxidative decarboxylation of pyruvate to acetyl-CoA. In mammals, PDHc is tightly regulated by phosphorylation-dephosphorylation of three serine residues in the thiamin-dependent pyruvate dehydrogenase (E1) component. In vivo, inactivation of human PDHc correlates mostly with phosphorylation of serine 264, which is located at the entrance of the substrate channel leading to the active site of E1. Despite intense investigations, the molecular mechanism of this inactivation has remained enigmatic. Here, a detailed analysis of microscopic steps of catalysis in human wild-type PDHc-E1 and pseudophosphorylation variant Ser264Glu elucidates how phosphorylation of Ser264 affects catalysis. Whereas the intrinsic reactivity of the active site in catalysis of pyruvate decarboxylation remains nearly unaltered, the preceding binding of substrate to the enzyme's active site via the substrate channel and the subsequent reductive acetylation of the E2 component are severely slowed in the phosphorylation variant. The structure of pseudophosphorylation variant Ser264Glu determined by X-ray crystallography reveals no differences in the three-dimensional architecture of the phosphorylation loop or of the active site, when compared to those of the wild-type enzyme. However, the channel leading to the active site is partially obstructed by the side chain of residue 264 in the variant. By analogy, a similar obstruction of the substrate channel can be anticipated to result from a phosphorylation of Ser264. The kinetic and thermodynamic results in conjunction with the structure of Ser264Glu suggest that phosphorylation blocks access to the active site by imposing a steric and electrostatic barrier for substrate binding and active site coupling with the E2 component. As a Ser264Gln variant, which carries no charge at position 264, is also selectively deficient in pyruvate binding and reductive acetylation of E2, we conclude that mostly steric effects account for inhibition of PDHc by phosphorylation.
- Institut für Biochemie und Biotechnologie, Martin-Luther-Universität Halle-Wittenberg, Kurt-Mothes-Strasse 3, 06120 Halle/Saale, Germany.
Organizational Affiliation: