Novel bis(indolyl)maleimide pyridinophanes that are potent, selective inhibitors of glycogen synthase kinase-3.
Zhang, H.C., Bonaga, L.V., Ye, H., Derian, C.K., Damiano, B.P., Maryanoff, B.E.(2007) Bioorg Med Chem Lett 17: 2863-2868
- PubMed: 17350261 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.02.059
- Primary Citation Related Structures:
2OW3 - PubMed Abstract:
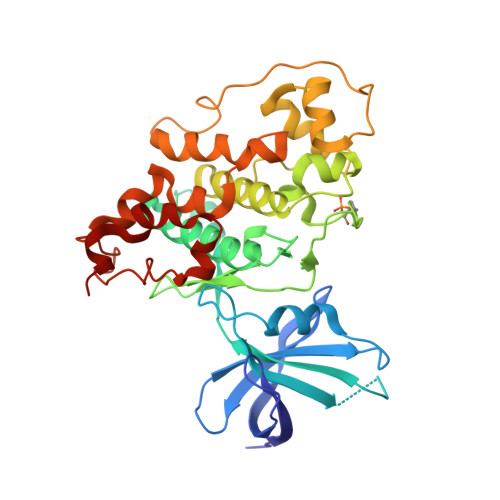
Novel bis(indolyl)maleimide pyridinophanes 3, 9a, 9b, 10a, 10b, and 11 were prepared by cobalt-mediated [2+2+2] cycloaddition of an appropriate alpha,omega-diyne with an N,N-dialkylcyanamide. These macrocyclic heterophanes were found to be potent, selective inhibitors of glycogen synthase kinase-3beta. An X-ray structure of a co-crystal of GSK-3beta and 3 (IC(50)=3nM) depicts the hydrogen bonding and hydrophobic interactions in the ATP-binding pocket of this serine/threonine protein kinase.
- Vascular Research Team, Johnson & Johnson Pharmaceutical Research & Development, Spring House, PA 19477-0776, USA. hzhang@prdus.jnj.com
Organizational Affiliation: