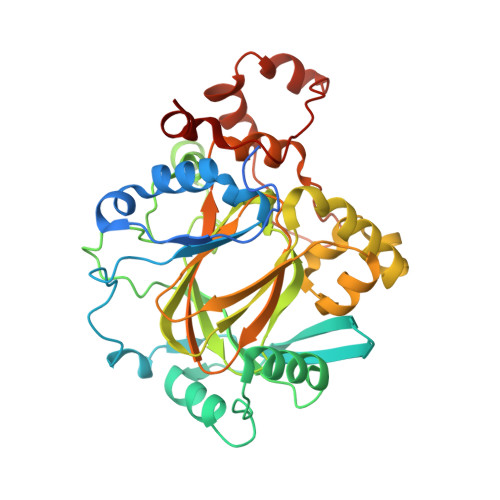
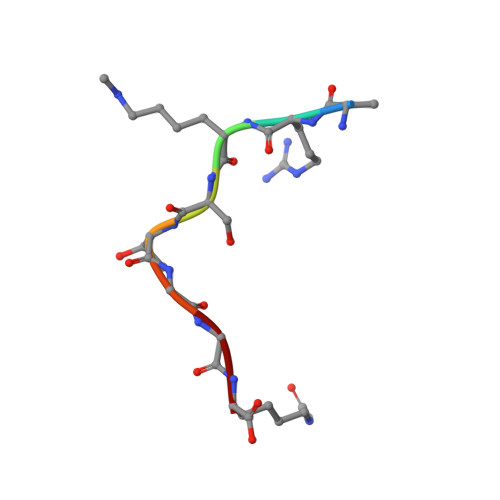
Crystal structures of histone demethylase JMJD2A reveal basis for substrate specificity.
Ng, S.S., Kavanagh, K.L., McDonough, M.A., Butler, D., Pilka, E.S., Lienard, B.M., Bray, J.E., Savitsky, P., Gileadi, O., von Delft, F., Rose, N.R., Offer, J., Scheinost, J.C., Borowski, T., Sundstrom, M., Schofield, C.J., Oppermann, U.(2007) Nature 448: 87-91
- PubMed: 17589501 Search on PubMed
- DOI: https://doi.org/10.1038/nature05971
- Primary Citation Related Structures:
2OQ6, 2OQ7, 2OS2, 2OT7, 2OX0 - PubMed Abstract:
Post-translational histone modification has a fundamental role in chromatin biology and is proposed to constitute a 'histone code' in epigenetic regulation. Differential methylation of histone H3 and H4 lysyl residues regulates processes including heterochromatin formation, X-chromosome inactivation, genome imprinting, DNA repair and transcriptional regulation. The discovery of lysyl demethylases using flavin (amine oxidases) or Fe(II) and 2-oxoglutarate as cofactors (2OG oxygenases) has changed the view of methylation as a stable epigenetic marker. However, little is known about how the demethylases are selective for particular lysyl-containing sequences in specific methylation states, a key to understanding their functions. Here we reveal how human JMJD2A (jumonji domain containing 2A), which is selective towards tri- and dimethylated histone H3 lysyl residues 9 and 36 (H3K9me3/me2 and H3K36me3/me2), discriminates between methylation states and achieves sequence selectivity for H3K9. We report structures of JMJD2A-Ni(II)-Zn(II) inhibitor complexes bound to tri-, di- and monomethyl forms of H3K9 and the trimethyl form of H3K36. The structures reveal a lysyl-binding pocket in which substrates are bound in distinct bent conformations involving the Zn-binding site. We propose a mechanism for achieving methylation state selectivity involving the orientation of the substrate methyl groups towards a ferryl intermediate. The results suggest distinct recognition mechanisms in different demethylase subfamilies and provide a starting point to develop chemical tools for drug discovery and to study and dissect the complexity of reversible histone methylation and its role in chromatin biology.
- Structural Genomics Consortium, Botnar Research Center, University of Oxford, Oxford OX3 7LD, UK.
Organizational Affiliation: