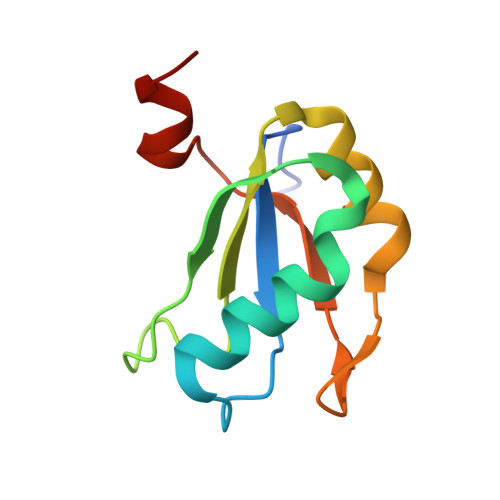
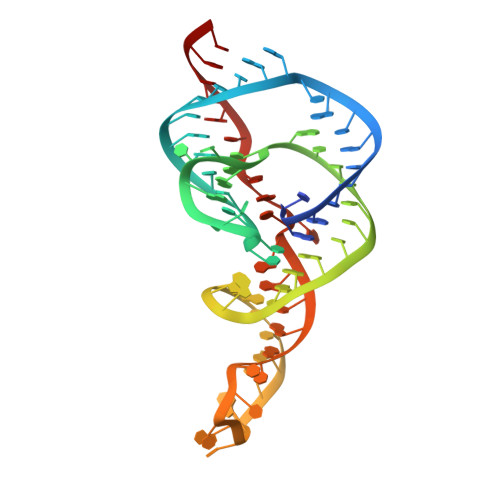
Structural roles of monovalent cations in the HDV ribozyme.
Ke, A., Ding, F., Batchelor, J.D., Doudna, J.A.(2007) Structure 15: 281-287
- PubMed: 17355864 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2007.01.017
- Primary Citation Related Structures:
2OIH, 2OJ3 - PubMed Abstract:
The hepatitis delta virus (HDV) ribozyme catalyzes viral RNA self-cleavage through general acid-base chemistry in which an active-site cytidine and at least one metal ion are involved. Monovalent metal ions support slow catalysis and were proposed to substitute for structural, but not catalytic, divalent metal ions in the RNA. To investigate the role of monovalent cations in ribozyme structure and function, we determined the crystal structure of the precursor HDV ribozyme in the presence of thallium ions (Tl(+)). Two Tl(+) ions can occupy a previously observed divalent metal ion hexahydrate-binding site located near the scissile phosphate, but are easily competed away by cobalt hexammine, a magnesium hexahydrate mimic and potent reaction inhibitor. Intriguingly, a third Tl(+) ion forms direct inner-sphere contacts with the ribose 2'-OH nucleophile and the pro-S(p) scissile phosphate oxygen. We discuss possible structural and catalytic implications of monovalent cation binding for the HDV ribozyme mechanism.
- Department of Molecular and Cell Biology, University of California at Berkeley, Berkeley, CA 94720, USA.
Organizational Affiliation: