Discovery of the HCV NS3/4A protease inhibitor (1R,5S)-N-[3-amino-1-(cyclobutylmethyl)-2,3-dioxopropyl]-3- [2(S)-[[[(1,1-dimethylethyl)amino]carbonyl]amino]-3,3-dimethyl-1-oxobutyl]- 6,6-dimethyl-3-azabicyclo[3.1.0]hexan-2(S)-carboxamide (Sch 503034) II. Key steps in structure-based optimization
Prongay, A.J., Guo, Z., Yao, N., Pichardo, J., Fischmann, T., Strickland, C., Myers, J., Weber, P.C., Beyer, B.M., Ingram, R., Hong, Z., Prosise, W.W., Ramanathan, L., Taremi, S.S., Yarosh-Tomaine, T., Zhang, R., Senior, M., Yang, R.S., Malcolm, B., Arasappan, A., Bennett, F., Bogen, S.L., Chen, K., Jao, E., Liu, Y.T., Lovey, R.G., Saksena, A.K., Venkatraman, S., Girijavallabhan, V., Njoroge, F.G., Madison, V.(2007) J Med Chem 50: 2310-2318
- PubMed: 17444623 Search on PubMed
- DOI: https://doi.org/10.1021/jm060173k
- Primary Citation Related Structures:
2O8M, 2OBO, 2OBQ, 2OC0, 2OC1, 2OC7, 2OC8 - PubMed Abstract:
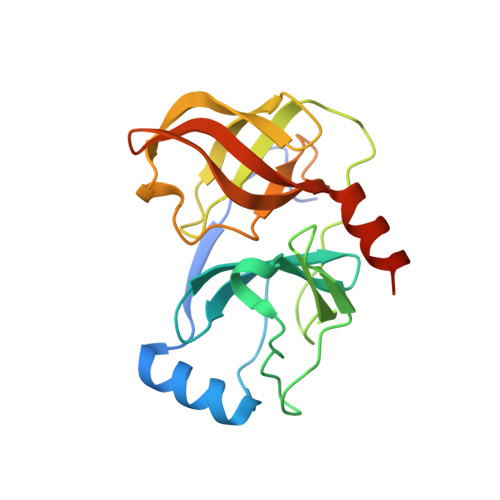
The structures of both the native holo-HCV NS3/4A protease domain and the protease domain with a serine 139 to alanine (S139A) mutation were solved to high resolution. Subsequently, structures were determined for a series of ketoamide inhibitors in complex with the protease. The changes in the inhibitor potency were correlated with changes in the buried surface area upon binding the inhibitor to the active site. The largest contribution to the binding energy arises from the hydrophobic interactions of the P1 and P2 groups as they bind to the S1 and S2 pockets [the numbering of the subsites is as defined in Berger, A.; Schechter, I. Philos. Trans. R. Soc. London, Ser. B 1970, 257, 249-264]. This correlation of the changes in potency with increased buried surface area contributed directly to the design of a potent tripeptide inhibitor of the HCV NS3/4A protease that is currently in clinical trials.
- Schering-Plough Research Institute, 2015 Galloping Hill Road, Kenilworth, New Jersey 07033, USA. andrew.prongay@spcorp.com
Organizational Affiliation: